|
|
Did you know...
Since version 1.5.0.5 PyMOL comes with a Plugin Manager, which can be used to load plugins such as those in the PyMOL Script Repo.
Features
- Install/Uninstall plugins
- Disable plugins and load them on demand to optimize PyMOL's startup time
- Configure the plugin search path
Appending additional paths to the Plugin Manager
Should your scripts be located in several alternative locations, it is possible to append additional directories to the Plugin Manager.
- Plugin > Plugin Manager > Settings > Add new directory...
Screenshots
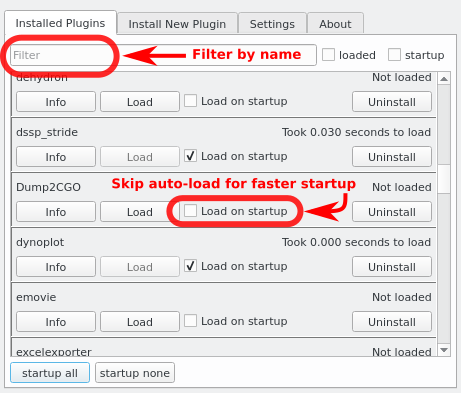
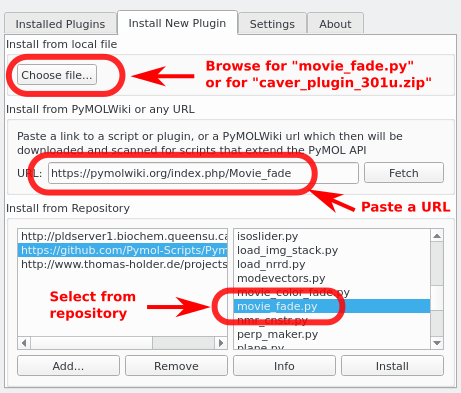
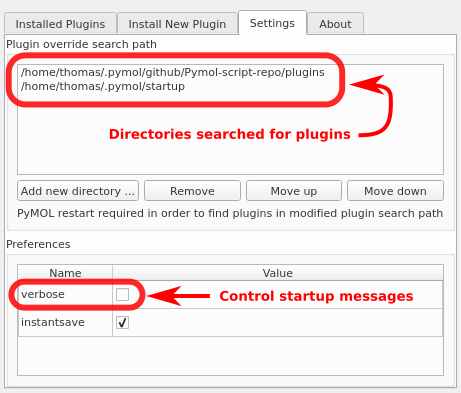
See Also
|
|
|
 A Random PyMOL-generated Cover. See Covers.
|



