Findseq
Appearance
| Type | Python Script |
|---|---|
| Download | scripts/findseq.py |
| Author(s) | Jason Vertrees |
| License | BSD |
| This code has been put under version control in the project Pymol-script-repo | |
Overview & Motivation
Anyone ever give you a protein and then say, find the sequence "FLVEW"? Well, this script will find any string or regular expression in a given object and return that selection for you. Here's an example,
reinitialize
import findseq
# fetch two sugar-binding PDB
fetch 1tvn, async=0
# Now, find FATEW in 1tvn, similarly
findseq FATEW, 1tvn
# lower-case works, too
findseq fatew, 1tvn
# how about a regular expression?
findseq F.*W, 1tvn
# Find the regular expression:
# ..H[TA]LVWH
# in the few proteins loaded.
# I then showed them as sticks and colored them to highlight matched AAs
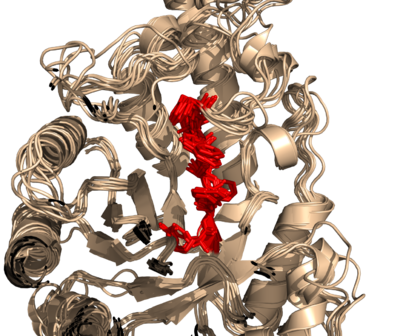
for x in cmd.get_names(): findseq.findseq("..H[TA]LVWH", x, "sele_"+x, firstOnly=1)
Usage
I built this to be rather flexible. You call it as:
findseq needle, haystack[, selName[, het[, firstOnly ]]]
where the options are:
- needle the sequence of amino acids to find. Should be a string of one letter amino acid abbreviations. Can also be a string-style regular expression (eg. FW.*QQ).
- haystack the PyMOL object or selection in which to search
- selName the name of the returned selection. If you leave this blank, it'll be foundSeqXYZ where XYZ is some random integer (eg. foundSeq1435); if you supply sele then the usual PyMOL (sele) is used; and, finally, if it's anything else, then that will be used verbatim. Defaults to foundSeqXYZ so as not to overwrite any selections you might have in sele.
- het 0/1 -- if 0 then heteroatoms are not considered; if 1 then they are; defaults to 0
- firstOnly 0/1 -- if 0 then all matches are selected and returned; if 1 then only the first is returned
See Also
select_pepseq@Psico