Difference between revisions of "Split states"
Jump to navigation
Jump to search
m |
m |
||
| Line 7: | Line 7: | ||
<source lang="python"> | <source lang="python"> | ||
load fileName.pdb1, name | load fileName.pdb1, name | ||
| − | + | split_states name | |
delete name | delete name | ||
</source> | </source> | ||
| Line 15: | Line 15: | ||
<source lang="python"> | <source lang="python"> | ||
load 1vls.pdb1, 1vls | load 1vls.pdb1, 1vls | ||
| − | + | split_states 1vls | |
dele 1vls | dele 1vls | ||
</source> | </source> | ||
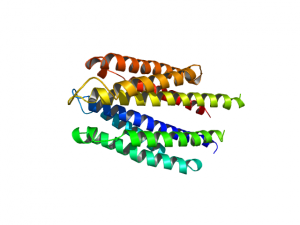
[[Image:1vls1.png|thumb|left|1VLS as a monomer. This is the state of 1VLS when I load the molecule (and select cartoon representation).]] | [[Image:1vls1.png|thumb|left|1VLS as a monomer. This is the state of 1VLS when I load the molecule (and select cartoon representation).]] | ||
[[Image:1vls1_dimer.png|left|thumb|1VLS as a dimer using the split_states command.]] | [[Image:1vls1_dimer.png|left|thumb|1VLS as a dimer using the split_states command.]] | ||
Revision as of 07:31, 3 August 2005
Overview
Split_States splits and orients multiple models and multimers from the biological unit file.
Using
To use split_states simply Load your molecule
load fileName.pdb1, name
split_states name
delete name
Example
1VLS: A dimer.
load 1vls.pdb1, 1vls
split_states 1vls
dele 1vls