File list
Jump to navigation
Jump to search
This special page shows all uploaded files.
| Date | Name | Thumbnail | Size | User | Description | Versions |
|---|---|---|---|---|---|---|
| 22:50, 8 March 2014 | Surface ramp above mode.jpg (file) |  |
163 KB | Speleo3 | illustration of the surface_ramp_above_mode/surface_solvent setting combinations | 1 |
| 11:02, 4 February 2014 | Amyloid.png (file) |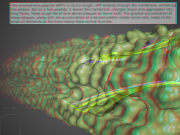  |
1.77 MB | Inchoate | Example (simulated) amyloid beta. | 1 |
| 10:59, 4 February 2014 | 3D-B-DNA.png (file) |  |
1.43 MB | Inchoate | Anaglyph image of DNA | 1 |
| 11:48, 17 December 2013 | Ana4 focal ray.png (file) |  |
659 KB | Inchoate | Ray trace with anaglyph and focal blur. | 1 |
| 12:20, 18 November 2013 | Bondlab.png (file) |  |
198 KB | SebastianRaschka | 1 | |
| 12:20, 18 November 2013 | Bondvis.png (file) |  |
255 KB | SebastianRaschka | 1 | |
| 12:19, 18 November 2013 | Hydrobond action.png (file) |  |
187 KB | SebastianRaschka | 1 | |
| 12:19, 18 November 2013 | Bondinfo.png (file) |  |
154 KB | SebastianRaschka | 1 | |
| 12:19, 18 November 2013 | Add hydrogens.png (file) | 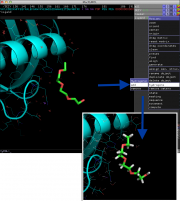 |
212 KB | SebastianRaschka | 1 | |
| 20:15, 6 October 2013 | Optimize.png (file) | 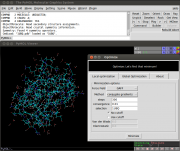 |
163 KB | OsvaldoMartin | optimize plugin GUI | 1 |
| 17:20, 3 October 2013 | Wrappers histogram.png (file) | 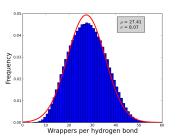 |
42 KB | OsvaldoMartin | Histogram of the number of wrappers per hydrongen bond for a set of high-quality X-ray protein structures. | 1 |
| 08:08, 10 September 2013 | Molelogo.png (file) |  |
8 KB | LukasPravda | Mole 2.0 logo | 1 |
| 07:46, 10 September 2013 | 1jj2.png (file) | 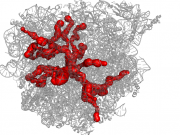 |
953 KB | LukasPravda | Example of channel system of large ribosomal subunit (1JJ2) | 1 |
| 05:12, 2 September 2013 | Example 1 FAD 1C0L.png (file) | 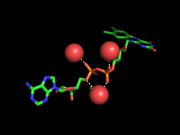 |
261 KB | GianlucaTomasello | 1 | |
| 18:25, 28 August 2013 | Cheshift.png (file) |  |
133 KB | OsvaldoMartin | The new file reflects the new color code used in CheShift-2 | 2 |
| 07:57, 29 July 2013 | Figure2a.png (file) |  |
129 KB | TomaszMakarewicz | Main Window in Dynamics PyMOL Plugin | 1 |
| 13:50, 24 July 2013 | Pairwise3.png (file) | 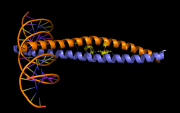 |
204 KB | PietroGattiLafranconi | 1 | |
| 13:50, 24 July 2013 | Pairwise2.png (file) | 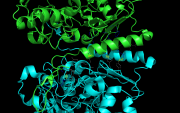 |
397 KB | PietroGattiLafranconi | 1 | |
| 13:49, 24 July 2013 | Pairwise1.png (file) | 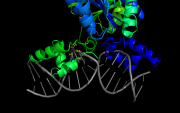 |
261 KB | PietroGattiLafranconi | 1 | |
| 03:41, 9 July 2013 | MtsslTrilaterate screenshot.png (file) |  |
317 KB | Gha | 1 | |
| 10:35, 4 July 2013 | MtsslDock results.jpg (file) |  |
265 KB | Gha | 1 | |
| 10:33, 4 July 2013 | MtsslDock distanceTable.jpg (file) |  |
80 KB | Gha | 1 | |
| 10:31, 4 July 2013 | MtsslDock Import Labels.jpg (file) |  |
91 KB | Gha | 1 | |
| 10:15, 4 July 2013 | MtsslDock screenshot.jpg (file) |  |
329 KB | Gha | 1 | |
| 15:53, 21 May 2013 | Heatmap-CMPyMOL.PNG (file) |  |
46 KB | Venky | 1 | |
| 16:37, 20 May 2013 | CMPyMOL.PNG (file) | 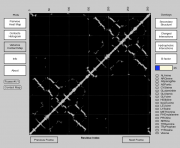 |
84 KB | Venky | 1 | |
| 09:12, 17 May 2013 | Cover Paunescu.jpg (file) |  |
2.22 MB | Inchoate | Paunescu's Cover | 1 |
| 13:17, 1 May 2013 | Contact density.png (file) |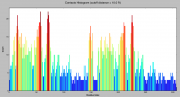  |
38 KB | Venky | 1 | |
| 13:17, 1 May 2013 | Heatmap.png (file) | 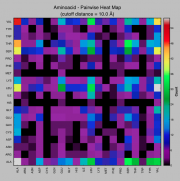 |
53 KB | Venky | 1 | |
| 14:31, 30 April 2013 | CMPyMOL-Screenshot.PNG (file) | 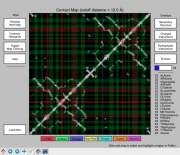 |
201 KB | Venky | Screenshot of CMPyMOL software that interfaces with PyMOL to combine 3D visualization of protein and 2D contact map representation. | 1 |
| 11:56, 26 April 2013 | 20091111 Structure cover.jpg (file) |  |
130 KB | Jaredsampson | Cover of the 11 November 2009 issue of Structure. Artwork by Jared Sampson, Xiangpeng Kong lab, New York University School of Medicine. | 1 |
| 04:53, 23 April 2013 | Cgo arrow example.png (file) | 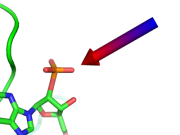 |
34 KB | Speleo3 | CGO arrow, made with cgo_arrow script | 1 |
| 10:55, 17 April 2013 | StylizedFocalBlur.png (file) | 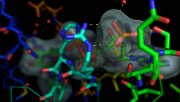 |
1.32 MB | Inchoate | 1 | |
| 12:07, 12 April 2013 | Stylized bns.png (file) | 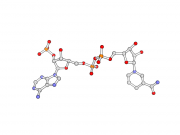 |
65 KB | Inchoate | Stylized ball and sticks | 1 |
| 21:54, 7 March 2013 | It.png (file) | 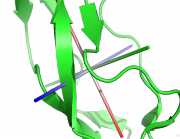 |
157 KB | Inchoate | inertia tensor | 1 |
| 12:09, 31 January 2013 | MtsslWizardv1-1.jpg (file) |  |
282 KB | Gha | 1 | |
| 11:17, 13 January 2013 | DistancesRH ex3.png (file) | 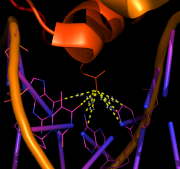 |
190 KB | PietroGattiLafranconi | 1 | |
| 11:16, 13 January 2013 | DistancesRH ex2.png (file) |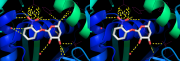  |
262 KB | PietroGattiLafranconi | 1 | |
| 11:16, 13 January 2013 | DistancesRH ex1.png (file) | 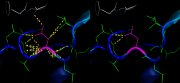 |
271 KB | PietroGattiLafranconi | 1 | |
| 01:13, 3 January 2013 | ParM.gif (file) |  |
82 KB | Inchoate | 1 | |
| 04:47, 6 December 2012 | Connect cutoff.png (file) | 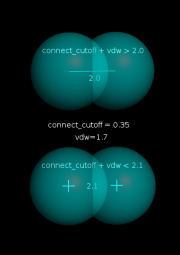 |
32 KB | Speleo3 | connect_cutoff setting example | 1 |
| 10:47, 20 September 2012 | Nb spheres.png (file) | 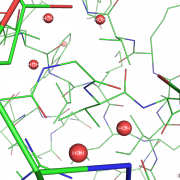 |
186 KB | Hongbo zhu | This png is generated using PyMOL 1.5.0.1. PDB structure model 1acb.pdb is rendered. | 1 |
| 06:53, 15 September 2012 | Proteins 08010 c1 sp Ob.png (file) |  |
264 KB | LarsS | Proteins Cover October 2012 - MD simulations of GroEL | 1 |
| 02:32, 18 August 2012 | Shader bug fixed.png (file) | 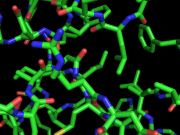 |
183 KB | TakanoriNakane | Corrected version of 'shader_bug.png' | 1 |
| 02:31, 18 August 2012 | Shader bug.png (file) | 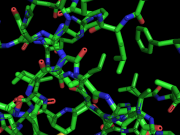 |
202 KB | TakanoriNakane | Cylinder shader bug which can be fixed by 'set cylinder_shader_ff_workaround, on' | 1 |
| 09:16, 26 June 2012 | Poseviewscreenshot.png (file) | 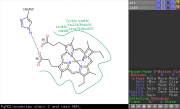 |
54 KB | Speleo3 | screenshot for poseview example: fetch 1a00, async=0 poseview chain C and resn HEM | 1 |
| 16:22, 12 June 2012 | GLSLToonCartoon.png (file) |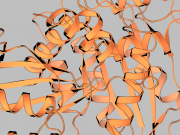  |
484 KB | Shiven | 1 | |
| 16:22, 12 June 2012 | GLSLToonSperes.png (file) |  |
400 KB | Shiven | 1 | |
| 14:45, 8 June 2012 | Stanfield.png (file) | 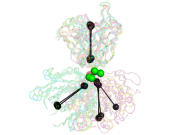 |
199 KB | Jaredsampson | Example PDBs from Stanfield, et al., JMB 2006 as shown in Fig. 1 of that paper. elbow_angle.py was used to calculate the elbow angles and draw a representation of the vectors used in the calculation. | 1 |
| 14:29, 8 June 2012 | Elbow angle 3ghe.png (file) | 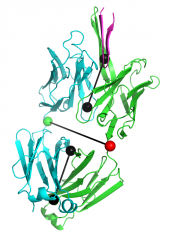 |
191 KB | Jaredsampson | View of the elbow_angle vector representation for the Fab fragment from PDB 3ghe. | 1 |