Cealign plugin
Update
Yeah! The C code that plugs into PyMol has been completed. It's a little slower than the plain C++ code I wrote, but that's what you get when passing data from PyMol to Python to C, fiddle with it, pass it back to Python to PyMol for some more quick math. The alignment times for the two proteins mentioned below (1B50 and 1C0M) on my machine with the new C module is about 1-3 second (with a full CPU load for other intensive tasks running in the background; this shows great improvement over the pure Python alignment times). Once the code is cleaned up (and I'm not too embarrassed to post it) and some bugs are worked out, I'll post it. The current bugs are:
- Some alignments don't center right
- Missing residues cause problems
- Memory leaks galore, I'm sure
The code consists of:
- qkabsch.py
- cealign.py
- ccealignmodule.c
- ccealignmodule.h
- setup.py
Also, I provide the option of aligning based solely upon RMSD or upon the better CE-Score. See the References for information on the CE Score.
Introduction
This script is a Python implementation of the CE algorithm pioneered by Drs. Shindyalov and Bourne (See References). It is a fast, accurate structure-based protein alignment algorithm. There are a few changes from the original code (See Notes), and "fast" depends on your machine and the implementation. That is, on my machine --- a relatively fast 64-bit machine --- I can align two 400+ amino acid structures in about 0.300 s with the C++ implementation. In Python however, two 165 amino acid proteins took about 35 seconds!
When coupled to the Kabsch algorithm, this should be able to align any two protein structures, using just the alpha carbon coordinates.
This plugs into PyMol very easily. See The Code and The Examples for installation and usage.
Documentation is forthcoming.
Comparison to PyMol
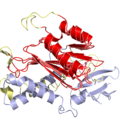
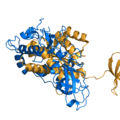
PyMol's structure alignment algorithm is fast and robust. However, its first step is to perform a sequence alignment of the two selections. Thus, proteins in the twilight zone or those having a low sequence identity, may not align well. Because CE is a structure-based alignment, this is not a problem. Look at the following example. The image at LEFT was the result of CE-aligning two proteins (1C0M to 1BCO). The result is 88 aligned (alpha carbons) residues (not atoms) at 2.78 Angstroms. The image on the RIGHT shows the results from PyMol's align command: an alignment of 221 atoms (not residues) at an RMSD of 15.7 Angstroms. To make the alignment easier to see, cealign (actually the Kabsch code) colors the aligned residues differently.
Notes
- The Python implementation is slow. This is most likely due to the fact that I'm not a very good Python coder. This is the initial version; if you can improve it, got for it. That's what open source is all about.
- This implementation requires the Kabsch algorithm I wrote to do the optimal superposition of the two structures once the residue pairings are determined.
- This implementation also uses the "CE-score" which is a statistically determined score that performs more reliably than does RMSD. I also provide the RMSD if you don't like the CE-score.
- I deviate from the original publication in that I use Kabsch's algorithm to align the two structures; nothing iterative.
- I deviate from Kabsch's algorithm by using the SVD solution, which is fast, accurate and easy to code (in comparison to the original elegant proof).
- This code is essentially a poor-man's translation of my C++ code.
- I deliberately left out the final optimization step (wiggling gaps on high scoring alignments) from the original paper. It is not relevant for my project. Someone else will have to code that.
Installation
Requirements
- Numpy
- Python 2.4+
Directions
- uncompress the distribution file cealign-VERSION.tgz
- cd cealign-VERSION
- sudo python setup.py install
- insert "run DIR_TO_CEALIGN/cealign.py" into your .pymolrc file
- load some molecules
- run, cealign molecule1, molecule2
- enjoy
The Code
Coming soon. (It works, but I'm trying to find ways to speed it up in Python.)
Examples
cealign 1cll, 1ggz
cealign 1kao, 1ctq
cealign 1fao, 1eaz
References
Text taken from PubMed and formatted for the wiki. The first reference is the most important for this code.
- Shindyalov IN, Bourne PE. Protein structure alignment by incremental combinatorial extension (CE) of the optimal path. Protein Eng. 1998 Sep;11(9):739-47. PMID: 9796821 [PubMed - indexed for MEDLINE]
- Jia Y, Dewey TG, Shindyalov IN, Bourne PE. A new scoring function and associated statistical significance for structure alignment by CE. J Comput Biol. 2004;11(5):787-99. PMID: 15700402 [PubMed - indexed for MEDLINE]
- Pekurovsky D, Shindyalov IN, Bourne PE. A case study of high-throughput biological data processing on parallel platforms. Bioinformatics. 2004 Aug 12;20(12):1940-7. Epub 2004 Mar 25. PMID: 15044237 [PubMed - indexed for MEDLINE]
- Shindyalov IN, Bourne PE. An alternative view of protein fold space. Proteins. 2000 Feb 15;38(3):247-60. PMID: 10713986 [PubMed - indexed for MEDLINE]