Uploads by Inchoate
Jump to navigation
Jump to search
This special page shows all uploaded files.
| Date | Name | Thumbnail | Size | Description | Versions |
|---|---|---|---|---|---|
| 15:25, 22 April 2011 | Selection overaly1.png (file) |  |
21 KB | selection indicators overlay atoms | 1 |
| 15:24, 22 April 2011 | Selection overaly0.png (file) |  |
21 KB | selection overlays atoms (off) | 1 |
| 12:59, 11 April 2011 | Bg grad.png (file) |  |
153 KB | background gradient | 1 |
| 12:56, 11 April 2011 | Screen shot 2011-04-11 at 12.55.49 PM.png (file) |  |
153 KB | Example image with the bg_gradient set. | 1 |
| 02:12, 8 April 2011 | Shade.png (file) | 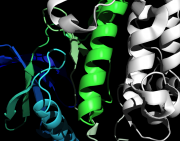 |
319 KB | ray_direct_shade on | 1 |
| 02:11, 8 April 2011 | No shade.png (file) | 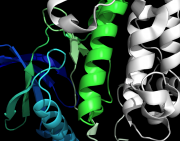 |
363 KB | ray_direct_shade off | 1 |
| 02:04, 8 April 2011 | Ray blend red.png (file) | 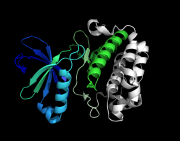 |
227 KB | red blended out | 1 |
| 02:03, 8 April 2011 | Ray blend off.png (file) | 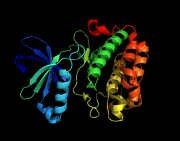 |
240 KB | ray blend off | 1 |
| 02:47, 5 March 2011 | Rnt.png (file) | 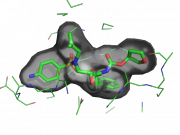 |
132 KB | Ray-normal transparency example | 1 |
| 23:33, 2 March 2011 | Highlight ss1.png (file) | 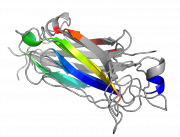 |
236 KB | example of common highlighted secondary structures | 1 |
| 15:43, 15 February 2011 | Salam.jpg (file) |  |
147 KB | mol. micro. Volume 79, Feb, 2011 | 1 |
| 16:06, 8 February 2011 | Map set ex.png (file) | 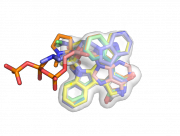 |
229 KB | Example consensus volume of the aligned ligands. | 1 |
| 10:39, 1 February 2011 | Apbs ex.png (file) |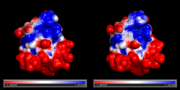  |
164 KB | Side-by-side map comparisons | 1 |
| 12:51, 2 December 2010 | Anaglyph1.png (file) | 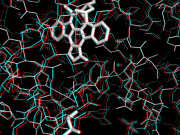 |
346 KB | 1 | |
| 23:27, 19 October 2010 | 1rmv fiber.png (file) | 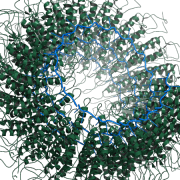 |
427 KB | 1RMV fiber, showing BiologicalUnit at work. | 1 |
| 11:32, 29 September 2010 | PyMOLreferenceCard-test.zip (file) | 68 KB | test | 1 | |
| 17:49, 28 September 2010 | Rtg10.png (file) | 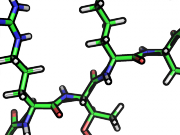 |
94 KB | zoomed in and ray trace gain, 10 | 1 |
| 17:49, 28 September 2010 | Rtg0.png (file) | 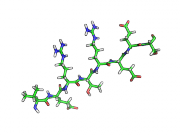 |
74 KB | ray trace gain, 0 | 1 |
| 17:49, 28 September 2010 | Rtgd.png (file) | 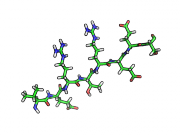 |
78 KB | ray trace gain, default | 1 |
| 17:45, 28 September 2010 | Rsch1.png (file) |  |
60 KB | ribbon side chain helper on | 1 |
| 17:45, 28 September 2010 | Rsch0.png (file) |  |
74 KB | Ribbon side chain helper off | 1 |
| 18:19, 15 September 2010 | Surface residues ex.png (file) |  |
666 KB | 1 | |
| 09:20, 13 September 2010 | Helix orientation2.png (file) |  |
76 KB | helix angle. | 1 |
| 09:15, 13 September 2010 | Helix orientation1.png (file) |  |
67 KB | example helix orientation | 1 |
| 18:13, 28 June 2010 | Surface by prop3.png (file) |  |
244 KB | 1 | |
| 18:13, 28 June 2010 | Surface by prop2.png (file) |  |
217 KB | 1 | |
| 18:13, 28 June 2010 | Surface by prop.png (file) |  |
546 KB | 1 | |
| 14:34, 28 June 2010 | BbPlane1.png (file) | 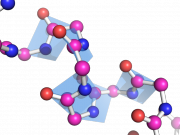 |
164 KB | bbplane ex1 | 1 |
| 14:33, 28 June 2010 | BbPlane2.png (file) | 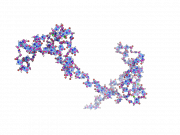 |
184 KB | bbplane ex 2 | 1 |
| 14:33, 28 June 2010 | BbPlane3.png (file) | 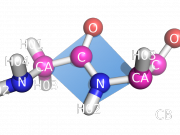 |
138 KB | bbplane ex. 3 | 1 |
| 10:17, 9 June 2010 | Tilt shift.png (file) | 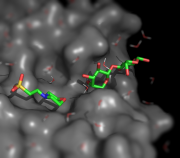 |
197 KB | Example of simulating tilt-shift. | 1 |
| 23:38, 15 April 2010 | After.png (file) | 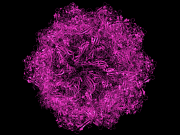 |
405 KB | After manual symexp. | 1 |
| 23:37, 15 April 2010 | Before.png (file) |  |
18 KB | Before manual symexp. | 1 |
| 19:34, 31 March 2010 | Sel orig.png (file) |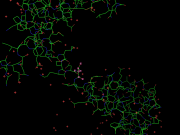  |
93 KB | Original selection. | 1 |
| 19:34, 31 March 2010 | Sel hi.png (file) |  |
72 KB | 1 | |
| 19:32, 31 March 2010 | Sel lo.png (file) |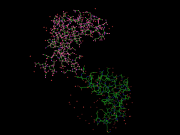  |
73 KB | 1 | |
| 22:10, 24 March 2010 | Map double3.png (file) | 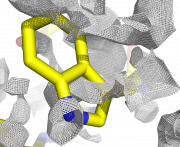 |
966 KB | 1 | |
| 22:09, 24 March 2010 | Map double2.png (file) | 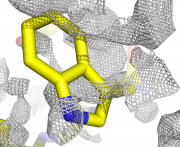 |
947 KB | 1 | |
| 22:08, 24 March 2010 | Map double.png (file) | 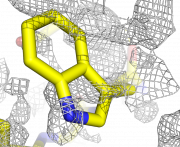 |
783 KB | 1 | |
| 22:07, 24 March 2010 | Map normal.png (file) |  |
582 KB | map at normal mesh | 1 |
| 10:53, 19 March 2010 | Gslike.png (file) |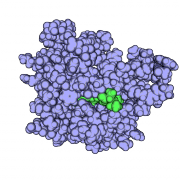  |
413 KB | 1 | |
| 00:39, 2 March 2010 | CircleR2.png (file) |  |
323 KB | Donut, width=150. | 1 |
| 00:33, 2 March 2010 | CircleR.png (file) |  |
211 KB | 1 | |
| 01:09, 13 February 2010 | Elines.png (file) |  |
1.14 MB | Electronic field lines. | 1 |
| 19:30, 16 January 2010 | Jbc mchu.jpeg (file) |  |
102 KB | Structure of the human mitotic checkpoint kinase Mps1 catalytic domain apo form (top) and inhibitor SP600125-bound form (bottom). "Crystal structure of the catalytic domain of the mitotic checkpoint kinase Mps1 in complex with SP600125. Chu ML, Chavas LM | 1 |
| 21:35, 12 January 2010 | Scm1.png (file) |  |
68 KB | Surface cavity 1 | 1 |
| 21:34, 12 January 2010 | Scm0.png (file) |  |
207 KB | Surface cavity 0 | 1 |
| 12:26, 19 November 2009 | JBC Apr18 2008.jpg (file) |  |
306 KB | Taylor AB, Hu G, Hart PJ, and McAlister-Henn L. (2008). Allosteric motions in structures of yeast NAD+-specific isocitrate dehydrogenase. J. Biol. Chem. 283,10872-10880. http://www.ncbi.nlm.nih.gov/pubmed/18256028?ordinalpos=1&itool=EntrezSystem2.PEntrez | 1 |
| 12:25, 19 November 2009 | ACR Jun 2007.jpg (file) |  |
90 KB | Quintanar L, Stoj C, Taylor AB, Hart PJ, Kosman DJ, and Solomon EI. (2007). Shall we dance? How a multicopper oxidase chooses its electron transfer partner. Acc. Chem. Res. 40, 445-452. http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=Retrieve& | 1 |
| 12:24, 19 November 2009 | ABB May1 2007.jpg (file) |  |
197 KB | Hu G, Taylor AB, McAlister-Henn L, and Hart PJ. (2007). Crystal structure of the yeast nicotinamidase Pnc1p. Arch. Biochem. Biophys. 461, 66-75. http://www.ncbi.nlm.nih.gov/entrez/query.fcgi?db=pubmed&cmd=Retrieve&dopt=AbstractPlus&list_uids=17382284&que | 1 |