FocalBlur: Difference between revisions
Jump to navigation
Jump to search
(angles as Fermat's spiral instead of random) |
(→Script) |
||
| Line 44: | Line 44: | ||
from shutil import rmtree | from shutil import rmtree | ||
from math import sin,cos,pi,sqrt | from math import sin,cos,pi,sqrt | ||
import subprocess | |||
def FocalBlur(aperture=2.0,samples=10,ray=0,width=0,height=0): | def FocalBlur(aperture=2.0,samples=10,ray=0,width=0,height=0): | ||
''' | ''' | ||
DESCRIPTION | DESCRIPTION | ||
Creates fancy figures by introducing a focal blur to the image. The object | Creates fancy figures by introducing a focal blur to the image. The object | ||
at the origin will be in focus. | at the origin will be in focus. | ||
AUTHOR | AUTHOR | ||
Jarl Underhaug | Jarl Underhaug | ||
University of Bergen | University of Bergen | ||
jarl_dot_underhaug_at_gmail_dot_com | jarl_dot_underhaug_at_gmail_dot_com | ||
Updates by Jason Vertrees and Thomas Holder | |||
USAGE | USAGE | ||
FocalBlur aperture=float, samples=int, ray=0/1, width=int, height=int | FocalBlur aperture=float, samples=int, ray=0/1, width=int, height=int | ||
EXAMPELS | EXAMPELS | ||
FocalBlur aperture=1, samples=100 | FocalBlur aperture=1, samples=100 | ||
FocalBlur aperture=2, samples=100, ray=1, width=600, height=400 | FocalBlur aperture=2, samples=100, ray=1, width=600, height=400 | ||
''' | ''' | ||
# Formalize the parameter types | |||
ray = (ray in ("True", "true", 1, "1")) | |||
aperture, samples = float(aperture), int(samples) | aperture, samples = float(aperture), int(samples) | ||
width, height = int(width), int(height) | |||
# Because of a bug, only custom sizes when raytracing | # Because of a bug, only custom sizes when raytracing | ||
if not ray: | #if not ray: | ||
# width=0 | |||
# height=0 | |||
# Create a temporary directory | # Create a temporary directory | ||
tmpdir = mkdtemp() | tmpdir = mkdtemp() | ||
# Get the orientation of the protein and the light | # Get the orientation of the protein and the light | ||
light = cmd.get('light')[1:-1] | light = cmd.get('light')[1:-1] | ||
light = [float(s) for s in light.split(',')] | light = [float(s) for s in light.split(',')] | ||
view = cmd.get_view() | view = cmd.get_view() | ||
# Rotate the protein and the light in order to create the blur | # Rotate the protein and the light in order to create the blur | ||
for frame in range(samples): | for frame in range(samples): | ||
| Line 93: | Line 98: | ||
xr = x/180.0*pi | xr = x/180.0*pi | ||
yr = y/180.0*pi | yr = y/180.0*pi | ||
# Rotate the protein | # Rotate the protein | ||
cmd.turn('x',x) | cmd.turn('x',x) | ||
cmd.turn('y',y) | cmd.turn('y',y) | ||
# Rotate the light | # Rotate the light | ||
ly = light[1]*cos(xr)-light[2]*sin(xr) | ly = light[1]*cos(xr)-light[2]*sin(xr) | ||
| Line 104: | Line 109: | ||
lz = lz*cos(yr)-lx*sin(yr) | lz = lz*cos(yr)-lx*sin(yr) | ||
cmd.set('light',[lx,ly,lz]) | cmd.set('light',[lx,ly,lz]) | ||
curFile = "%s/frame-%04d.png" % (tmpdir,frame) | |||
# Save the image | # Save the image | ||
if ray: | |||
cmd.png(curFile,width=width,height=height,ray=ray,quiet=1) | |||
else: | |||
cmd.png(curFile,quiet=1) | |||
# Return the protein and the light to the original orientation | # Return the protein and the light to the original orientation | ||
cmd.set('light',light) | cmd.set('light',light) | ||
cmd.set_view(view) | cmd.set_view(view) | ||
# Create a blured image of all the frames | # Create a blured image of all the frames | ||
r = subprocess.call('convert %s/frame-*.png +matte -average %s/blur.png' % (tmpdir,tmpdir),shell=True) | |||
# Load the blured image | |||
print "load %s/blur.png" % tmpdir | |||
cmd.load('%s/blur.png' % tmpdir) | |||
# Delete the temporary files | # Delete the temporary files | ||
rmtree(tmpdir) | rmtree(tmpdir) | ||
cmd.extend('FocalBlur', FocalBlur) | cmd.extend('FocalBlur', FocalBlur) | ||
Revision as of 04:04, 9 June 2011
Description
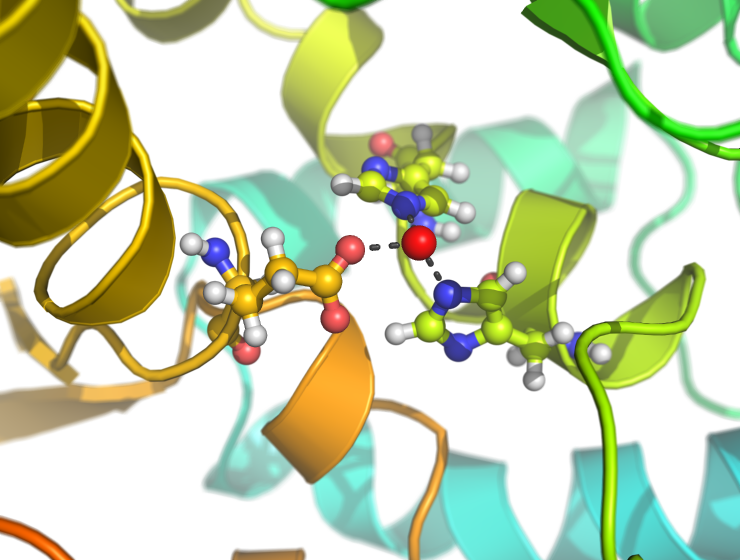
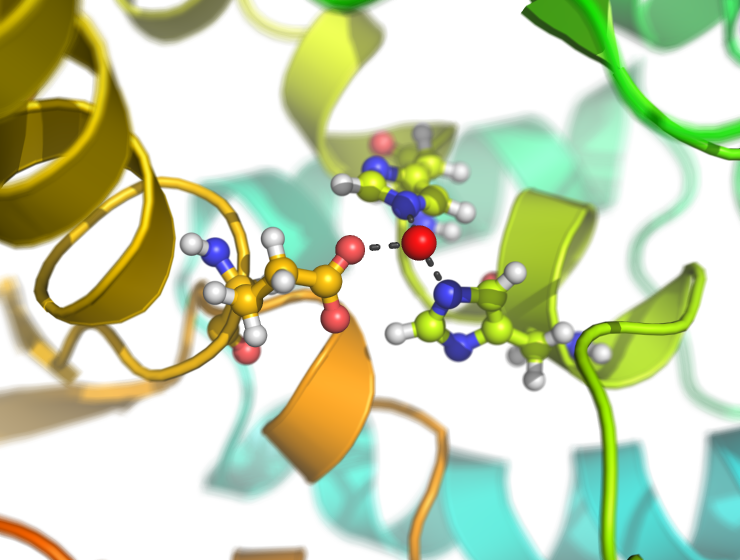
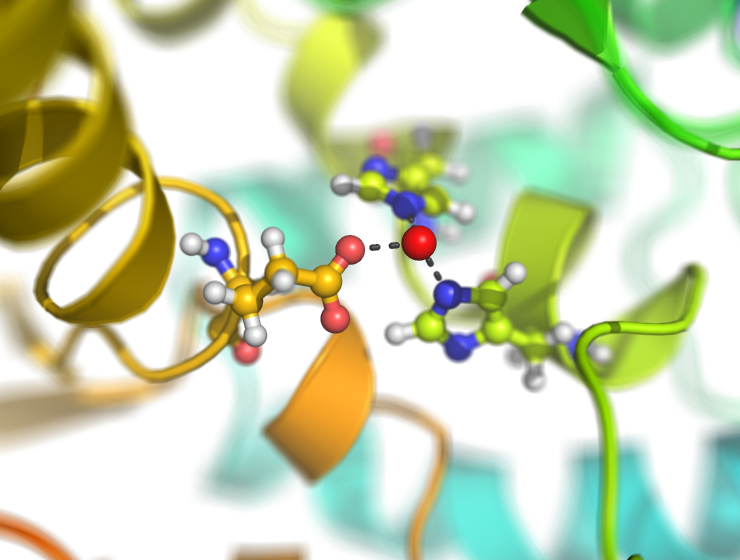
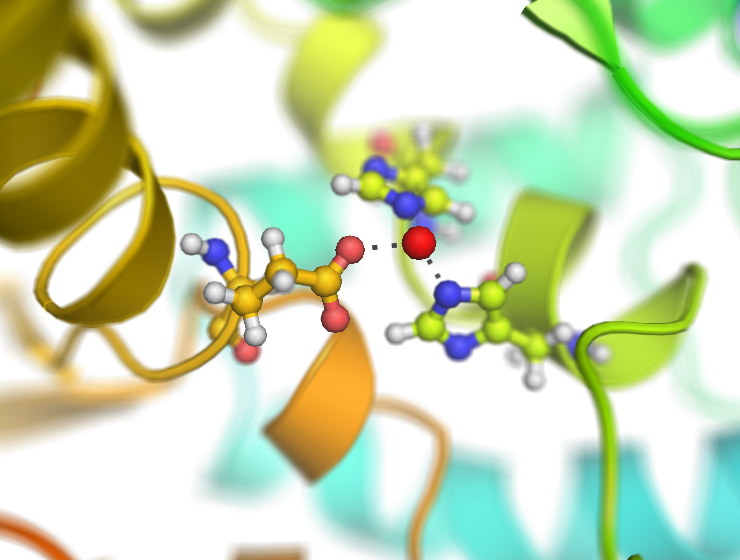
This script creates fancy figures by introducing a focal blur to the image. The object at the origin will be in focus.
Usage
Load the script using the run command. Execute the script using PyMOL syntax:
FocalBlur aperture=2.0,samples=100,ray=1
or using python syntax:
FocalBlur(aperture=2.0,samples=100,ray=1)
For additional options, see the script comments.
Notes
- When using raytracing, the image creation will take n times longer than normal, where n is the number of samples.
- The script uses ImageMagick for creating the blured image. It has only been tested on Linux
- The aperture is a purely arbitrary value and not related to f stops on a camera.
- There is a bug preventing custom image sizes when not using raytracing.
Examples
Script
Load the script using the run command
from pymol import cmd
from os import system
from tempfile import mkdtemp
from shutil import rmtree
from math import sin,cos,pi,sqrt
import subprocess
def FocalBlur(aperture=2.0,samples=10,ray=0,width=0,height=0):
'''
DESCRIPTION
Creates fancy figures by introducing a focal blur to the image. The object
at the origin will be in focus.
AUTHOR
Jarl Underhaug
University of Bergen
jarl_dot_underhaug_at_gmail_dot_com
Updates by Jason Vertrees and Thomas Holder
USAGE
FocalBlur aperture=float, samples=int, ray=0/1, width=int, height=int
EXAMPELS
FocalBlur aperture=1, samples=100
FocalBlur aperture=2, samples=100, ray=1, width=600, height=400
'''
# Formalize the parameter types
ray = (ray in ("True", "true", 1, "1"))
aperture, samples = float(aperture), int(samples)
width, height = int(width), int(height)
# Because of a bug, only custom sizes when raytracing
#if not ray:
# width=0
# height=0
# Create a temporary directory
tmpdir = mkdtemp()
# Get the orientation of the protein and the light
light = cmd.get('light')[1:-1]
light = [float(s) for s in light.split(',')]
view = cmd.get_view()
# Rotate the protein and the light in order to create the blur
for frame in range(samples):
# Angles to rotate protein and light
# Populate angles as Fermat's spiral
theta = frame * pi * 110.0/144.0
radius = 0.5 * aperture * sqrt(frame/float(samples-1))
x = cos(theta) * radius
y = sin(theta) * radius
xr = x/180.0*pi
yr = y/180.0*pi
# Rotate the protein
cmd.turn('x',x)
cmd.turn('y',y)
# Rotate the light
ly = light[1]*cos(xr)-light[2]*sin(xr)
lz = light[2]*cos(xr)+light[1]*sin(xr)
lx = light[0]*cos(yr)+lz*sin(yr)
lz = lz*cos(yr)-lx*sin(yr)
cmd.set('light',[lx,ly,lz])
curFile = "%s/frame-%04d.png" % (tmpdir,frame)
# Save the image
if ray:
cmd.png(curFile,width=width,height=height,ray=ray,quiet=1)
else:
cmd.png(curFile,quiet=1)
# Return the protein and the light to the original orientation
cmd.set('light',light)
cmd.set_view(view)
# Create a blured image of all the frames
r = subprocess.call('convert %s/frame-*.png +matte -average %s/blur.png' % (tmpdir,tmpdir),shell=True)
# Load the blured image
print "load %s/blur.png" % tmpdir
cmd.load('%s/blur.png' % tmpdir)
# Delete the temporary files
rmtree(tmpdir)
cmd.extend('FocalBlur', FocalBlur)