AAindex: Difference between revisions
Jump to navigation
Jump to search
(added figure) |
(automatic download) |
||
| Line 3: | Line 3: | ||
AAindex is a database of numerical indices representing various physicochemical and biochemical properties of amino acids and pairs of amino acids. See http://www.genome.jp/aaindex/ | AAindex is a database of numerical indices representing various physicochemical and biochemical properties of amino acids and pairs of amino acids. See http://www.genome.jp/aaindex/ | ||
This script is a python parser for the AAindex flat files which | This script is a python parser for the AAindex flat files which will be downloaded from ftp://ftp.genome.jp/pub/db/community/aaindex/ | ||
The script provides two PyMOL commands (but can also be used without PyMOL). | The script provides two PyMOL commands (but can also be used without PyMOL). | ||
| Line 36: | Line 32: | ||
<source lang="python"> | <source lang="python"> | ||
import aaindex | import aaindex | ||
aaindex2b KYTJ820101 | aaindex2b KYTJ820101 | ||
spectrum b, yellow_white_blue | spectrum b, yellow_white_blue | ||
| Line 46: | Line 40: | ||
<source lang="python"> | <source lang="python"> | ||
''' | ''' | ||
(c) 2010 Thomas Holder | (c) 2010-2011 Thomas Holder, MPI for Developmental Biology | ||
Python parser for AAindex: Amino Acid Index Database | Python parser for AAindex: Amino Acid Index Database | ||
| Line 56: | Line 50: | ||
pmf | pmf | ||
''' | ''' | ||
import sys, os | |||
_aaindex = dict() | _aaindex = dict() | ||
| Line 106: | Line 102: | ||
assert aaj is None | assert aaj is None | ||
return self.index.get(aai, d) | return self.index.get(aai, d) | ||
def __getitem__(self, aai): | |||
return self.get(aai) | |||
def median(self): | def median(self): | ||
x = sorted(filter(None, self.index.values())) | x = sorted(filter(None, self.index.values())) | ||
| Line 140: | Line 138: | ||
except: | except: | ||
return d | return d | ||
def __getitem__(self, aaij): | |||
return self.get(aaij[0], aaij[1]) | |||
def median(self): | def median(self): | ||
x = [] | x = [] | ||
| Line 155: | Line 155: | ||
if len(_aaindex) == 0: | if len(_aaindex) == 0: | ||
init() | init() | ||
return _aaindex | return _aaindex[key] | ||
def _float_or_None(x): | def _float_or_None(x): | ||
| Line 162: | Line 162: | ||
return float(x) | return float(x) | ||
def init(path=None, index=' | def init(path=None, index='13'): | ||
''' | ''' | ||
Read in the aaindex files. You need to run this (once) before you can | Read in the aaindex files. You need to run this (once) before you can | ||
| Line 169: | Line 169: | ||
aaindex files are read in. | aaindex files are read in. | ||
''' | ''' | ||
index = str(index) | |||
if path is None: | if path is None: | ||
for path in [os.path.split(__file__)[0], '.', cmd.get('fetch_path')]: | |||
if os.path.exists(os.path.join(path, 'aaindex' + index[0])): | |||
break | |||
print >> sys.stderr, 'path =', path | print >> sys.stderr, 'path =', path | ||
if '1' in index: | if '1' in index: | ||
_parse(path + '/aaindex1', Record) | _parse(path + '/aaindex1', Record) | ||
| Line 191: | Line 191: | ||
`MarixRecord` for aaindex2 and aaindex3. | `MarixRecord` for aaindex2 and aaindex3. | ||
''' | ''' | ||
if not os.path.exists(filename): | |||
import urllib | |||
url = 'ftp://ftp.genome.jp/pub/db/community/aaindex/' + os.path.split(filename)[1] | |||
print 'Downloading "%s"' % (url) | |||
filename = urllib.urlretrieve(url, filename)[0] | |||
print 'Saved to "%s"' % (filename) | |||
f = open(filename) | |||
current = rec() | current = rec() | ||
| Line 359: | Line 360: | ||
EXAMPLES | EXAMPLES | ||
# distance dependent c-beta contact potentials | # distance dependent c-beta contact potentials | ||
| Line 398: | Line 396: | ||
print '%s %.1f-%.1f' % (key[i], cutoff[i], cutoff[i+1]) | print '%s %.1f-%.1f' % (key[i], cutoff[i], cutoff[i+1]) | ||
idmap = dict() | |||
cmd.iterate_state(state, '(%s) or (%s)' % (selection1, selection2), | cmd.iterate_state(state, '(%s) or (%s)' % (selection1, selection2), | ||
' | 'idmap[model,index] = [(resn,name),(x,y,z)]', space={'idmap': idmap}) | ||
twoN = cmd.count_atoms(selection1) + cmd.count_atoms(selection2) | twoN = cmd.count_atoms(selection1) + cmd.count_atoms(selection2) | ||
pairs = cmd.find_pairs(selection1, selection2, cutoff=max(cutoff), | pairs = cmd.find_pairs(selection1, selection2, cutoff=max(cutoff), | ||
| Line 416: | Line 414: | ||
count = 0 | count = 0 | ||
for id1, id2 in pairs: | for id1, id2 in pairs: | ||
a1 = | a1 = idmap[id1] | ||
a2 = | a2 = idmap[id2] | ||
r = cpv.distance(a1[1], a2[1]) | r = cpv.distance(a1[1], a2[1]) | ||
for i in i_list: | for i in i_list: | ||
Revision as of 14:55, 4 April 2011
AAindex is a database of numerical indices representing various physicochemical and biochemical properties of amino acids and pairs of amino acids. See http://www.genome.jp/aaindex/
This script is a python parser for the AAindex flat files which will be downloaded from ftp://ftp.genome.jp/pub/db/community/aaindex/
The script provides two PyMOL commands (but can also be used without PyMOL).
- aaindex2b: Loads numerical indices from aaindex1 as b-factors into your structure
- pmf: Potential of Mean Force (aaindex3)
Python Example
Consider the script is called aaindex.py, it is placed somewhere in your PYTHONPATH and the aaindex flatfiles are found in the current directory.
import aaindex
aaindex.init(path='.')
aaindex.grep('volume')
x = aaindex.get('KRIW790103')
print x
print x.get('A')
aaindex.grep('blosum')
x = aaindex.get('HENS920102')
print x.get('A', 'K')
PyMOL Example
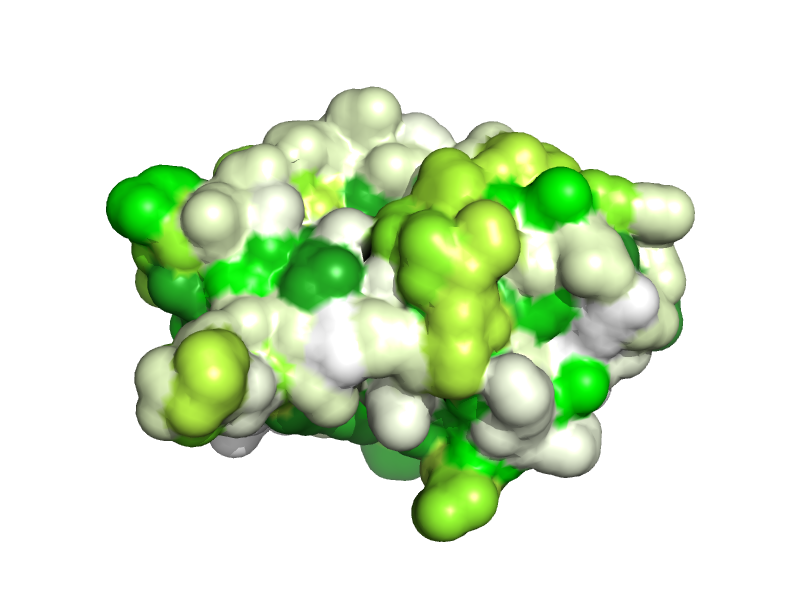
import aaindex
aaindex2b KYTJ820101
spectrum b, yellow_white_blue
show surface
The Script
'''
(c) 2010-2011 Thomas Holder, MPI for Developmental Biology
Python parser for AAindex: Amino Acid Index Database
http://www.genome.jp/aaindex/
PyMOL commands:
aaindex2b
pmf
'''
import sys, os
_aaindex = dict()
_pymol_auto_arg_update = lambda: None
def search(pattern, searchtitle=True, casesensitive=False):
'''
Search for pattern in description and title (optional) of all records and
return matched records as list. By default search case insensitive.
'''
whatcase = lambda i: i
if not casesensitive:
pattern = pattern.lower()
whatcase = lambda i: i.lower()
matches = []
for record in _aaindex.itervalues():
if pattern in whatcase(record.desc) or searchtitle and pattern in whatcase(record.title):
matches.append(record)
return matches
def grep(pattern):
'''
Search for pattern in title and description of all records (case
insensitive) and print results on standard output.
'''
for record in search(pattern):
print record
class Record:
'''
Amino acid index (AAindex) Record
'''
aakeys = 'ARNDCQEGHILKMFPSTWYV'
def __init__(self):
self.key = None
self.desc = ''
self.ref = ''
self.authors = ''
self.title = ''
self.journal = ''
self.correlated = dict()
self.index = dict()
self.comment = ''
def extend(self, row):
i = len(self.index)
for x in row:
self.index[self.aakeys[i]] = x
i += 1
def get(self, aai, aaj=None, d=None):
assert aaj is None
return self.index.get(aai, d)
def __getitem__(self, aai):
return self.get(aai)
def median(self):
x = sorted(filter(None, self.index.values()))
half = len(x)/2
if len(x) % 2 == 1:
return x[half]
return (x[half-1] + x[half])/2.0
def __str__(self):
desc = self.desc.replace('\n', ' ').strip()
return '%s(%s: %s)' % (self.__class__.__name__, self.key, desc)
class MatrixRecord(Record):
'''
Matrix record for mutation matrices or pair-wise contact potentials
'''
def __init__(self):
Record.__init__(self)
self.index = []
self.rows = dict()
self.cols = dict()
def extend(self, row):
self.index.append(row)
def _get(self, aai, aaj):
i = self.rows[aai]
j = self.cols[aaj]
return self.index[i][j]
def get(self, aai, aaj, d=None):
try:
return self._get(aai, aaj)
except:
pass
try:
return self._get(aaj, aai)
except:
return d
def __getitem__(self, aaij):
return self.get(aaij[0], aaij[1])
def median(self):
x = []
for y in self.index:
x.extend(filter(None, y))
x.sort()
if len(x) % 2 == 1:
return x[len(x)/2]
return sum(x[len(x)/2-1:len(x)/2+1])/2.0
def get(key):
'''
Get record for key
'''
if len(_aaindex) == 0:
init()
return _aaindex[key]
def _float_or_None(x):
if x == 'NA' or x == '-':
return None
return float(x)
def init(path=None, index='13'):
'''
Read in the aaindex files. You need to run this (once) before you can
access any records. If the files are not within the current directory,
you need to specify the correct directory path. By default all three
aaindex files are read in.
'''
index = str(index)
if path is None:
for path in [os.path.split(__file__)[0], '.', cmd.get('fetch_path')]:
if os.path.exists(os.path.join(path, 'aaindex' + index[0])):
break
print >> sys.stderr, 'path =', path
if '1' in index:
_parse(path + '/aaindex1', Record)
if '2' in index:
_parse(path + '/aaindex2', MatrixRecord)
if '3' in index:
_parse(path + '/aaindex3', MatrixRecord)
_pymol_auto_arg_update()
def init_from_file(filename, type=Record):
_parse(filename, type)
def _parse(filename, rec, quiet=True):
'''
Parse aaindex input file. `rec` must be `Record` for aaindex1 and
`MarixRecord` for aaindex2 and aaindex3.
'''
if not os.path.exists(filename):
import urllib
url = 'ftp://ftp.genome.jp/pub/db/community/aaindex/' + os.path.split(filename)[1]
print 'Downloading "%s"' % (url)
filename = urllib.urlretrieve(url, filename)[0]
print 'Saved to "%s"' % (filename)
f = open(filename)
current = rec()
lastkey = None
for line in f:
key = line[0:2]
if key[0] == ' ':
key = lastkey
if key == '//':
_aaindex[current.key] = current
current = rec()
elif key == 'H ':
current.key = line[2:].strip()
elif key == 'R ':
current.ref += line[2:]
elif key == 'D ':
current.desc += line[2:]
elif key == 'A ':
current.authors += line[2:]
elif key == 'T ':
current.title += line[2:]
elif key == 'J ':
current.journal += line[2:]
elif key == '* ':
current.comment += line[2:]
elif key == 'C ':
a = line[2:].split()
for i in range(0, len(a), 2):
current.correlated[a[i]] = float(a[i+1])
elif key == 'I ':
a = line[1:].split()
if a[0] != 'A/L':
current.extend(map(_float_or_None, a))
elif list(Record.aakeys) != [i[0] for i in a] + [i[-1] for i in a]:
print 'Warning: wrong amino acid sequence for', current.key
else:
try:
assert list(Record.aakeys[:10]) == [i[0] for i in a]
assert list(Record.aakeys[10:]) == [i[2] for i in a]
except:
print 'Warning: wrong amino acid sequence for', current.key
elif key =='M ':
a = line[2:].split()
if a[0] == 'rows':
if a[4] == 'rows':
a.pop(4)
assert a[3] == 'cols' and len(a) == 6
i = 0
for aa in a[2]:
current.rows[aa] = i
i += 1
i = 0
for aa in a[5]:
current.cols[aa] = i
i += 1
else:
current.extend(map(_float_or_None, a))
elif not quiet:
print 'Warning: line starts with "%s"' % (key)
lastkey = key
########## PYMOL ###########
# from Bio.SCOP.Raf import to_one_letter_code
# See also http://www.pymolwiki.org/index.php/Aa_codes
to_one_letter_code = {'PAQ': 'Y', 'AGM': 'R', 'ILE': 'I', 'PR3': 'C',
'GLN': 'Q', 'DVA': 'V', 'CCS': 'C', 'ACL': 'R', 'GLX': 'Z', 'GLY': 'G',
'GLZ': 'G', 'DTH': 'T', 'OAS': 'S', 'C6C': 'C', 'NEM': 'H', 'DLY': 'K',
'MIS': 'S', 'SMC': 'C', 'GLU': 'E', 'NEP': 'H', 'BCS': 'C', 'ASQ': 'D',
'ASP': 'D', 'SCY': 'C', 'SER': 'S', 'LYS': 'K', 'SAC': 'S', 'PRO': 'P',
'ASX': 'B', 'DGN': 'Q', 'DGL': 'E', 'MHS': 'H', 'ASB': 'D', 'ASA': 'D',
'NLE': 'L', 'DCY': 'C', 'ASK': 'D', 'GGL': 'E', 'STY': 'Y', 'SEL': 'S',
'CGU': 'E', 'ASN': 'N', 'ASL': 'D', 'LTR': 'W', 'DAR': 'R', 'VAL': 'V',
'CHG': 'A', 'TPO': 'T', 'CLE': 'L', 'GMA': 'E', 'HAC': 'A', 'AYA': 'A',
'THR': 'T', 'TIH': 'A', 'SVA': 'S', 'MVA': 'V', 'SAR': 'G', 'LYZ': 'K',
'BNN': 'A', '5HP': 'E', 'IIL': 'I', 'SHR': 'K', 'HAR': 'R', 'FME': 'M',
'PYX': 'C', 'ALO': 'T', 'PHI': 'F', 'ALM': 'A', 'PHL': 'F', 'MEN': 'N',
'TPQ': 'A', 'GSC': 'G', 'PHE': 'F', 'ALA': 'A', 'MAA': 'A', 'MET': 'M',
'UNK': 'X', 'LEU': 'L', 'ALY': 'K', 'SET': 'S', 'GL3': 'G', 'TRG': 'K',
'CXM': 'M', 'TYR': 'Y', 'SCS': 'C', 'DIL': 'I', 'TYQ': 'Y', '3AH': 'H',
'DPR': 'P', 'PRR': 'A', 'CME': 'C', 'IYR': 'Y', 'CY1': 'C', 'TYY': 'Y',
'HYP': 'P', 'DTY': 'Y', '2AS': 'D', 'DTR': 'W', 'FLA': 'A', 'DPN': 'F',
'DIV': 'V', 'PCA': 'E', 'MSE': 'M', 'MSA': 'G', 'AIB': 'A', 'CYS': 'C',
'NLP': 'L', 'CYQ': 'C', 'HIS': 'H', 'DLE': 'L', 'CEA': 'C', 'DAL': 'A',
'LLP': 'K', 'DAH': 'F', 'HMR': 'R', 'TRO': 'W', 'HIC': 'H', 'CYG': 'C',
'BMT': 'T', 'DAS': 'D', 'TYB': 'Y', 'BUC': 'C', 'PEC': 'C', 'BUG': 'L',
'CYM': 'C', 'NLN': 'L', 'CY3': 'C', 'HIP': 'H', 'CSO': 'C', 'TPL': 'W',
'LYM': 'K', 'DHI': 'H', 'MLE': 'L', 'CSD': 'A', 'HPQ': 'F', 'MPQ': 'G',
'LLY': 'K', 'DHA': 'A', 'DSN': 'S', 'SOC': 'C', 'CSX': 'C', 'OMT': 'M',
'DSP': 'D', 'PTR': 'Y', 'TRP': 'W', 'CSW': 'C', 'EFC': 'C', 'CSP': 'C',
'CSS': 'C', 'SCH': 'C', 'OCS': 'C', 'NMC': 'G', 'SEP': 'S', 'BHD': 'D',
'KCX': 'K', 'SHC': 'C', 'C5C': 'C', 'HTR': 'W', 'ARG': 'R', 'TYS': 'Y',
'ARM': 'R', 'DNP': 'A'}
def aaindex2b(key='KYTJ820101', selection='(all)', quiet=0, var='b'):
'''
DESCRIPTION
"aaindex" looks up the Amino Acid Index from
http://www.genome.jp/aaindex/
for the given key and assignes b-factors to the given selection. Unknown
residues get the average index value assigned.
USAGE
aaindex2b [key [, selection]]
ARGUMENTS
key = string: Key of AAindex entry
selection = string: atoms to assign b-factors {default: (all)}
EXAMPLE
# Hydropathy index by Kyte-Doolittle
aaindex2b KYTJ820101
spectrumany b, white yellow forest
show surface
'''
from pymol import cmd, stored
entry = get(key)
median = entry.median()
if int(quiet) != 0:
print entry.desc.strip()
def lookup(resn):
one_letter = to_one_letter_code.get(resn, 'X')
value = entry.get(one_letter)
if value is None:
return median
return value
stored.aaindex = lookup
cmd.alter(selection, var + '=stored.aaindex(resn)')
def pmf(key, cutoff=7.0, selection1='(name CB)', selection2='', state=1, quiet=1):
'''
DESCRIPTION
Potential of Mean Force
ARGUMENTS
key = string: aaindex key
cutoff = float: distance cutoff {default: 7.0}
cutoff = (float, float): distance shell
selection1 = string: atom selection {default: (name CB)}
selection2 = string: atom selection {default: selection1}
NOTES
Does also support a list of keys and a list of cutoffs to deal with
multiple distance shells.
EXAMPLES
# distance dependent c-beta contact potentials
pmf SIMK990101, 5, /2x19//A//CB
pmf SIMK990102, [5, 7.5], /2x19//A//CB
pmf [SIMK990101, SIMK990102, SIMK990103], [0, 5, 7.5, 10], /2x19//A//CB
# interface potential
sidechaincenters 2x19_scc, 2x19
pmf KESO980102, 7.0, /2x19_scc//A, /2x19_scc//B
distance /2x19_scc//A, /2x19_scc//B, cutoff=7.0
'''
from pymol import cmd, stored
from chempy import cpv
if cmd.is_string(key):
if key.lstrip().startswith('['):
key = cmd.safe_alpha_list_eval(key)
else:
key = [key]
if cmd.is_string(cutoff):
cutoff = eval(cutoff)
if not cmd.is_sequence(cutoff):
cutoff = [cutoff]
if len(cutoff) == len(key):
cutoff = [0.0] + list(cutoff)
if len(cutoff) != len(key) + 1:
print 'Error: Number of keys and number of cutoffs inconsistent'
return
state = int(state)
quiet = int(quiet)
if len(selection2) == 0:
selection2 = selection1
if not quiet and len(key) > 1:
print 'Distance shells:'
for i in range(len(key)):
print '%s %.1f-%.1f' % (key[i], cutoff[i], cutoff[i+1])
idmap = dict()
cmd.iterate_state(state, '(%s) or (%s)' % (selection1, selection2),
'idmap[model,index] = [(resn,name),(x,y,z)]', space={'idmap': idmap})
twoN = cmd.count_atoms(selection1) + cmd.count_atoms(selection2)
pairs = cmd.find_pairs(selection1, selection2, cutoff=max(cutoff),
state1=state, state2=state)
if len(pairs) == 0:
print 'Empty pair list'
return 0.0
matrix = map(get, key)
for i in matrix:
assert isinstance(i, MatrixRecord)
i_list = range(len(key))
u_sum = 0
count = 0
for id1, id2 in pairs:
a1 = idmap[id1]
a2 = idmap[id2]
r = cpv.distance(a1[1], a2[1])
for i in i_list:
if cutoff[i] <= r and r < cutoff[i+1]:
try:
aa1 = to_one_letter_code[a1[0][0]]
aa2 = to_one_letter_code[a2[0][0]]
u_sum += matrix[i].get(aa1, aa2)
count += 1
except:
print 'Failed for', a1[0], a2[0]
value = float(u_sum) / twoN
if not quiet:
print 'PMF: %.4f (%d contacts, %d residues)' % (value, count, twoN)
return value
try:
from pymol import cmd
cmd.extend('aaindex2b', aaindex2b)
cmd.extend('pmf', pmf)
def pymol_auto_arg_update():
aaindexkey_sc = cmd.Shortcut(_aaindex.keys())
cmd.auto_arg[0].update({
'aaindex2b' : [ aaindexkey_sc , 'aaindexkey' , ', ' ],
'pmf' : [ aaindexkey_sc , 'aaindexkey' , ', ' ],
})
cmd.auto_arg[1].update({
'aaindex2b' : [ cmd.selection_sc , 'selection' , '' ],
})
cmd.auto_arg[2].update({
'pmf' : [ cmd.selection_sc , 'selection' , '' ],
})
cmd.auto_arg[3].update({
'pmf' : [ cmd.selection_sc , 'selection' , '' ],
})
_pymol_auto_arg_update = pymol_auto_arg_update
except:
pass
# vi: ts=4:sw=4:smarttab:expandtab