Main Page: Difference between revisions
Jump to navigation
Jump to search
No edit summary |
No edit summary |
||
| Line 67: | Line 67: | ||
escapelinks=false | escapelinks=false | ||
openreferences=true | openreferences=true | ||
listseparators=[[,%PAGE%,|thumb|185px|A Random PyMOL-generated Cover. See [[Covers]].]], | listseparators=[[,%PAGE%,|thumb|185px|A Random PyMOL-generated Cover. See [[Covers]].]],\n | ||
</dpl> | </dpl> | ||
|} | |} | ||
Revision as of 14:09, 18 November 2009
| The community-run support site for the PyMOL molecular viewer. |
| Tutorials | Table of Contents | Commands |
| Script Library | Plugins | FAQ |
| Gallery | Covers | PyMOL Cheat Sheet (PDF) | GoogleSearch |
|
|
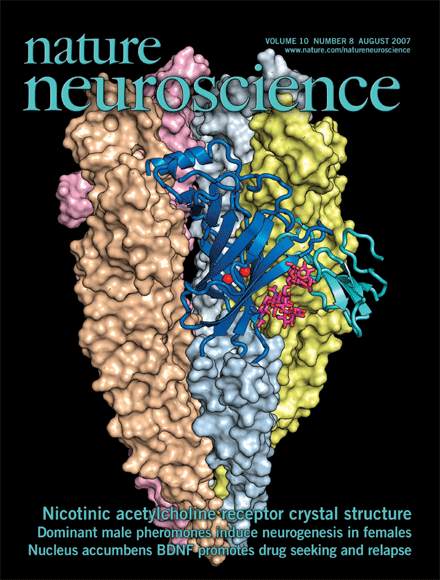 A Random PyMOL-generated Cover. See Covers.
| ||||||||||||||||||