All states: Difference between revisions
Jump to navigation
Jump to search
m (Added two images to illustrate the example.) |
|||
| Line 20: | Line 20: | ||
set all_states, on | set all_states, on | ||
</source> | </source> | ||
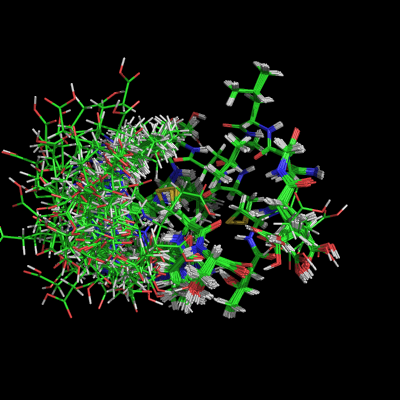
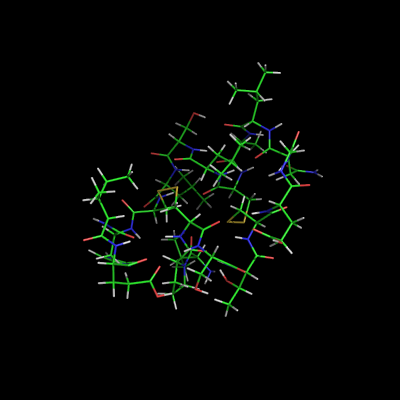
This shows the effect of turning on/off the '''all_states''' setting used with the script above. | |||
<gallery> | |||
Image:All_states_on.png|set all_states, on | |||
Image:All_states_off.png|set all_states, off | |||
</gallery> | |||
Revision as of 20:52, 14 September 2006
Overview
When set "on", this setting causes PyMOL to display all states or in NMR jargon: all the models in the ensemble. The 'default' behavior (OFF) can be overridden by placing the "set all_states, on" statement into your '.pymolrc' file, located in your login directory (under all flavors of unix).
Syntax
set all_states, on
set all_states, off
Example
import urllib2
pdbCode = '1BRV'
pdbUrl = 'http://www.rcsb.org/pdb/downloadFile.do?fileFormat=pdb&compression=NO&structureId='+pdbCode
pdbFile = urllib2.urlopen(pdbUrl)
pdbContent = pdbFile.read()
cmd.read_pdbstr(pdbContent, pdbCode)
set all_states, on
This shows the effect of turning on/off the all_states setting used with the script above.