Practical Pymol for Beginners: Difference between revisions
(added some images, still a work in progress...) |
(more text) |
||
| Line 1: | Line 1: | ||
Although PyMol has a powerful and flexible | Although PyMol has a powerful and flexible interface, it is complex, and can appear daunting to new users. This guide is intended to introduce the PyMol interface and basic tasks without leaving the mouse behind. | ||
== | ==The PyMol Interface== | ||
When PyMol is opened, two windows appear. The smaller window (called the "External GUI" in PyMol documentation) has the menu bar ('''File''', '''Edit''', '''Help''', '''Display''', etc), a set of buttons which are shortcuts to common commands, and the command line. | |||
[[Image:Viewer_guide.png|right|thumb|The PyMol Viewer Window]] | [[Image:Viewer_guide.png|right|thumb|The PyMol Viewer Window]] | ||
The PyMol | The second window is the PyMol viewer, which is where all the magic happens. In the viewer, 3D models are displayed, and the user interacts (eg rotates) and manipulates the model. The Object Control Panel is used to change colors, viewing mode, apply labels, remove objects, and just about anything else relating to objects. (An object is normally a group of molecules loaded from a PDB coordinate file.) Each object is displayed by name followed by a set of command buttons which control the object. Here are the menus and some of their options: | ||
* A - Actions: Close, etc | * A - Actions: Close, etc | ||
* S - Show: Change the way an object is represented, eg change stick view to cartoon view. | * S - Show: Change the way an object is represented, eg change stick view to cartoon view. | ||
| Line 14: | Line 11: | ||
* L - Label: Label atoms, residues, etc. | * L - Label: Label atoms, residues, etc. | ||
* C - Color: Change the color of atoms and groups. | * C - Color: Change the color of atoms and groups. | ||
The lower-left corner of the Viewer contains a guide to using the mouse, as well as a powerful selection tool. There is also another command line at the bottom of the Viewer ('''PyMOL>'''). | |||
==About the command line== | |||
The PyMol command line is a great tool that lets the experienced user change all sorts of options that simply don't appear in the point-and-click graphical interface. It can also be a lot faster. Combined with scripting, it is a powerful option for automating tasks and making intricate sets of changes. But, it's complex, and page upon page of PyMol documentation cover these commands, so we're going to ignore them as much as possible. | |||
Although this guide may include some text commands (''like this''), they're purely optional and meant to be informative. | |||
==Visualize a protein== | ==Visualize a protein== | ||
These instructions cover opening a PDB file and using PyMol as | These instructions cover opening a PDB file and using PyMol as tool for molecular exploration. | ||
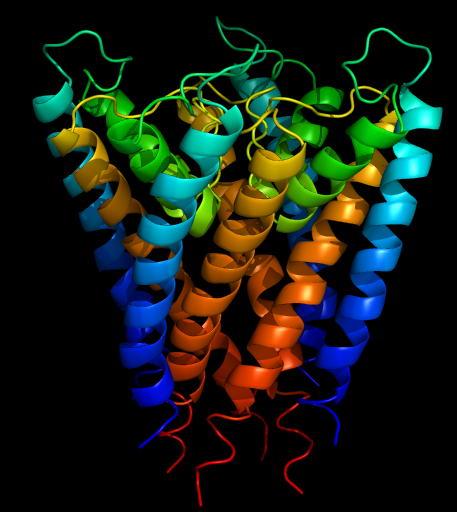
[[Image:Kchannel-rainbow.png|thumb| | [[Image:Kchannel-rainbow.png|thumb|right|The end result will look something like this]] | ||
# Obtain a PDB coordinates file for your favorite protein. (The [http://www.pdb.org/ | # Obtain a PDB coordinates file for your favorite protein. (The [http://www.pdb.org/ RCSB Protein Data Bank] is a public structure repository containing over 40,000 protein structures in PDB format available for download, not a bad place to look.) For this example, we're using the potassium channel from ''Streptomyces Lividans'' ([http://www.pdb.org/pdb/files/1bl8.pdb 1BL8]). | ||
# Open the PDB file using '''File''', '''Open...''' from the menu bar. The protein's structure will appear, probably rendered as | # Open the PDB file using '''File''', '''Open...''' from the menu bar. The protein's structure will appear, probably rendered as simple bonding lines. | ||
# | # The right side of the Viewer shows objects loaded from the PDB, and the command buttons. Each button contains a submenu that can be used to manipulate the object. Click '''S''', then '''cartoon''' to show the protein's secondary structure in popular cartoon form. | ||
# To change the color of each protein chain (as defined in the coordinate file), click '''C''' then select '''Chainbows''' from the ''' | #* Notice that the lines view is still visible on top of the cartoon view. To hide the lines, click '''H''' then '''lines'''. | ||
# Click and drag the protein to change the view. | # To change the color of each protein chain (as defined in the coordinate file), click '''C''' then select '''chainbows''' from the '''by chain''' menu. "Chainbows" colors residues in each protein chain as a rainbow that begins with blue and ends with red. | ||
#* Another common coloring method assigns a single color to each chain. Click '''C''' then select '''by chain''' from the '''by chain''' menu. | |||
# Click and drag the protein to change the view. Reminder: a legend for mouse actions is below the object control panel. | |||
#*(Cover mouse matrix abbreviations) | #*(Cover mouse matrix abbreviations) | ||
So, now what? Good question. PyMOL is a powerful program, and everyone uses it for something different. The remainder of this guide is devoted to common tasks that come in handy. | |||
==Saving an image== | |||
You've found the perfect view, and you'd like to save it? | |||
# Images are saved exactly as they appear in the viewer, including size. Change the window size and zoom in/out if it's not the proper size. | |||
#* For example, images for printing and presenting should be larger than images for a posting on a website. | |||
# Because PyMOL was designed to run on older computers, the maximum quality is not enabled by default. To change this, select '''Display''', '''Quality''', '''Maximum Quality'''. Notice that round things are rounder, curves are curvier, and color shading is smoother. | |||
# Once you've found the appropriate view, save an image using '''File''', '''Save Image...''' An picture of the current view will be saved in PNG format. | |||
'''Advanced:''' Using the ''[[ray]]'' command before saving will create a higher quality version of the molecule with shadows, etc. Depending on the size of the image and speed of the computer, it can take a while (seconds to minutes...), but the images created with ray tracing are usually spectacular. However, the ray tracing disappears if the view is changed in any way. | |||
==Getting unstuck== | |||
PyMOL is a program meant to be explored and played with, and sometimes funny things happen in the process. A few common scenarios: | |||
* ''The model disappeared:'' Sometimes while rotating and moving a model, it can get lost. Right-click the background of the viewer, and the ''Main Pop-Up'' should appear. Select '''reset''', and the model should return to view. | |||
* ''The model has funny colors, labels, etc and they won't go away:'' The '''H''' menu in the object control panel will remove unwanted details; however, sometimes it's difficult to know exactly what to remove. Select '''H''', then '''everything''' to hide all details. | |||
==Other== | ==Other== | ||
| Line 42: | Line 58: | ||
* Objects vs selection objects? | * Objects vs selection objects? | ||
* Pic of External GUI for command line | * Pic of External GUI for command line | ||
* s/PyMol/PyMOL | |||
* Dummy-style advanced notation. | |||
* Howto run commands... | |||
** Tone change. | |||
* intro and outro | |||
Revision as of 17:55, 10 April 2007
Although PyMol has a powerful and flexible interface, it is complex, and can appear daunting to new users. This guide is intended to introduce the PyMol interface and basic tasks without leaving the mouse behind.
The PyMol Interface
When PyMol is opened, two windows appear. The smaller window (called the "External GUI" in PyMol documentation) has the menu bar (File, Edit, Help, Display, etc), a set of buttons which are shortcuts to common commands, and the command line.
The second window is the PyMol viewer, which is where all the magic happens. In the viewer, 3D models are displayed, and the user interacts (eg rotates) and manipulates the model. The Object Control Panel is used to change colors, viewing mode, apply labels, remove objects, and just about anything else relating to objects. (An object is normally a group of molecules loaded from a PDB coordinate file.) Each object is displayed by name followed by a set of command buttons which control the object. Here are the menus and some of their options:
- A - Actions: Close, etc
- S - Show: Change the way an object is represented, eg change stick view to cartoon view.
- H - Hide: Things that are shown using S accumulate, and don't automatically replace the last view. H has commands for hiding unwanted representations.
- L - Label: Label atoms, residues, etc.
- C - Color: Change the color of atoms and groups.
The lower-left corner of the Viewer contains a guide to using the mouse, as well as a powerful selection tool. There is also another command line at the bottom of the Viewer (PyMOL>).
About the command line
The PyMol command line is a great tool that lets the experienced user change all sorts of options that simply don't appear in the point-and-click graphical interface. It can also be a lot faster. Combined with scripting, it is a powerful option for automating tasks and making intricate sets of changes. But, it's complex, and page upon page of PyMol documentation cover these commands, so we're going to ignore them as much as possible.
Although this guide may include some text commands (like this), they're purely optional and meant to be informative.
Visualize a protein
These instructions cover opening a PDB file and using PyMol as tool for molecular exploration.
- Obtain a PDB coordinates file for your favorite protein. (The RCSB Protein Data Bank is a public structure repository containing over 40,000 protein structures in PDB format available for download, not a bad place to look.) For this example, we're using the potassium channel from Streptomyces Lividans (1BL8).
- Open the PDB file using File, Open... from the menu bar. The protein's structure will appear, probably rendered as simple bonding lines.
- The right side of the Viewer shows objects loaded from the PDB, and the command buttons. Each button contains a submenu that can be used to manipulate the object. Click S, then cartoon to show the protein's secondary structure in popular cartoon form.
- Notice that the lines view is still visible on top of the cartoon view. To hide the lines, click H then lines.
- To change the color of each protein chain (as defined in the coordinate file), click C then select chainbows from the by chain menu. "Chainbows" colors residues in each protein chain as a rainbow that begins with blue and ends with red.
- Another common coloring method assigns a single color to each chain. Click C then select by chain from the by chain menu.
- Click and drag the protein to change the view. Reminder: a legend for mouse actions is below the object control panel.
- (Cover mouse matrix abbreviations)
So, now what? Good question. PyMOL is a powerful program, and everyone uses it for something different. The remainder of this guide is devoted to common tasks that come in handy.
Saving an image
You've found the perfect view, and you'd like to save it?
- Images are saved exactly as they appear in the viewer, including size. Change the window size and zoom in/out if it's not the proper size.
- For example, images for printing and presenting should be larger than images for a posting on a website.
- Because PyMOL was designed to run on older computers, the maximum quality is not enabled by default. To change this, select Display, Quality, Maximum Quality. Notice that round things are rounder, curves are curvier, and color shading is smoother.
- Once you've found the appropriate view, save an image using File, Save Image... An picture of the current view will be saved in PNG format.
Advanced: Using the ray command before saving will create a higher quality version of the molecule with shadows, etc. Depending on the size of the image and speed of the computer, it can take a while (seconds to minutes...), but the images created with ray tracing are usually spectacular. However, the ray tracing disappears if the view is changed in any way.
Getting unstuck
PyMOL is a program meant to be explored and played with, and sometimes funny things happen in the process. A few common scenarios:
- The model disappeared: Sometimes while rotating and moving a model, it can get lost. Right-click the background of the viewer, and the Main Pop-Up should appear. Select reset, and the model should return to view.
- The model has funny colors, labels, etc and they won't go away: The H menu in the object control panel will remove unwanted details; however, sometimes it's difficult to know exactly what to remove. Select H, then everything to hide all details.
Other
- Reinitialize hint
TODO:
- Capitalization for internal GUI features?
- Definition of an object?
- Objects vs selection objects?
- Pic of External GUI for command line
- s/PyMol/PyMOL
- Dummy-style advanced notation.
- Howto run commands...
- Tone change.
- intro and outro