BbPlane: Difference between revisions
Jump to navigation
Jump to search
m (→The Source) |
(multistate support, simplification of vertex ordering) |
||
| Line 32: | Line 32: | ||
from chempy import cpv | from chempy import cpv | ||
def bbPlane(objSel='(all)', color='white', transp=0.0): | def bbPlane(objSel='(all)', color='white', transp=0.0, state=1, name=None, quiet=1): | ||
""" | """ | ||
DESCRIPTION | DESCRIPTION | ||
| Line 45: | Line 45: | ||
transp = float: transparency component (0.0--1.0) {default: 0.0} | transp = float: transparency component (0.0--1.0) {default: 0.0} | ||
state = integer: object state, 0 for all states {default: 1} | |||
NOTES | NOTES | ||
| Line 53: | Line 55: | ||
# format input | # format input | ||
transp = float(transp) | transp = float(transp) | ||
state, quiet = int(state), int(quiet) | |||
if name is None: | |||
name = cmd.get_unused_name("backbonePlane") | |||
if state < 0: | |||
state = cmd.get_state() | |||
elif state == 0: | |||
for state in range(1, cmd.count_states(objSel)+1): | |||
bbPlane(objSel, color, transp, state, name, quiet) | |||
return | |||
stored.AAs = [] | stored.AAs = [] | ||
coords = dict() | coords = dict() | ||
| Line 62: | Line 75: | ||
for obj in cmd.get_object_list(objSel): | for obj in cmd.get_object_list(objSel): | ||
sel = obj + " and (" + objSel + ")" | sel = obj + " and (" + objSel + ")" | ||
for a in cmd.get_model(sel + " and n. CA").atom: | for a in cmd.get_model(sel + " and n. CA", state).atom: | ||
key = '/%s/%s/%s/%s' % (obj,a.segi,a.chain,a.resi) | key = '/%s/%s/%s/%s' % (obj,a.segi,a.chain,a.resi) | ||
stored.AAs.append(key) | stored.AAs.append(key) | ||
coords[key] = [a.coord,None,None] | coords[key] = [a.coord,None,None] | ||
for a in cmd.get_model(sel + " and n. O").atom: | for a in cmd.get_model(sel + " and n. O", state).atom: | ||
key = '/%s/%s/%s/%s' % (obj,a.segi,a.chain,a.resi) | key = '/%s/%s/%s/%s' % (obj,a.segi,a.chain,a.resi) | ||
if key in coords: | if key in coords: | ||
coords[key][1] = a.coord | coords[key][1] = a.coord | ||
for a in cmd.get_model("( | for a in cmd.get_model(sel + " and ((n. N extend 1 and e. H) or (r. PRO and n. CD))", state).atom: | ||
key = '/%s/%s/%s/%s' % (obj,a.segi,a.chain,a.resi) | key = '/%s/%s/%s/%s' % (obj,a.segi,a.chain,a.resi) | ||
if key in coords: | if key in coords: | ||
| Line 104: | Line 117: | ||
# need to order vertices to generate correct triangles for plane | # need to order vertices to generate correct triangles for plane | ||
# modified/added by B.Bell 8/18/2011 | # modified/added by B.Bell 8/18/2011 | ||
# modified by Thomas Holder 2012 | |||
if cpv.dot_product(cpv.sub(pos[0], pos[1]), cpv.sub(pos[2], pos[3])) < 0: | |||
vorder = [0,1,2,2,3,0] | |||
else: | |||
vorder = [0,1,2,3,2,1] | |||
# fill in the vertex data for the triangles; | # fill in the vertex data for the triangles; | ||
for i in vorder: | for i in vorder: | ||
| Line 127: | Line 132: | ||
# update the UI | # update the UI | ||
cmd.load_cgo(obj, name, state) | |||
cmd.load_cgo(obj, | cmd.set("cgo_transparency", transp, name) | ||
cmd.set("cgo_transparency", transp, | |||
cmd.extend("bbPlane", bbPlane) | cmd.extend("bbPlane", bbPlane) | ||
Revision as of 09:53, 26 January 2012
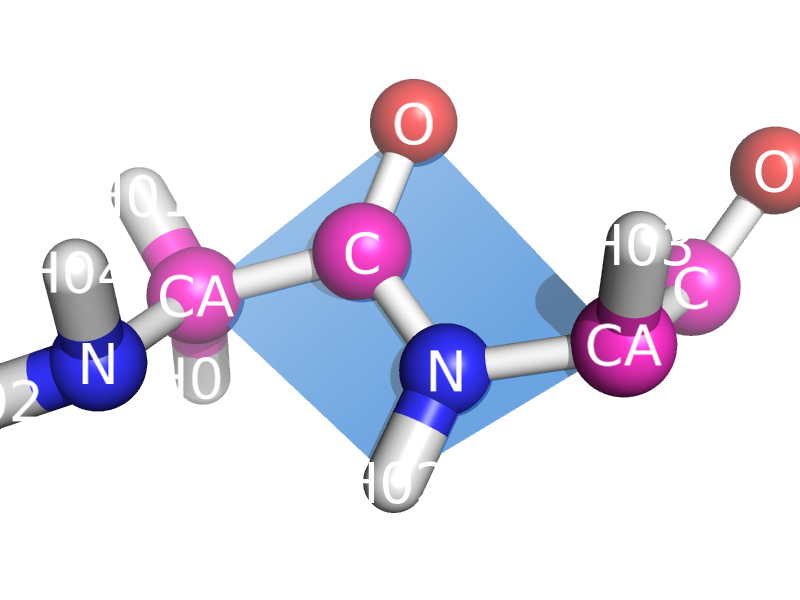
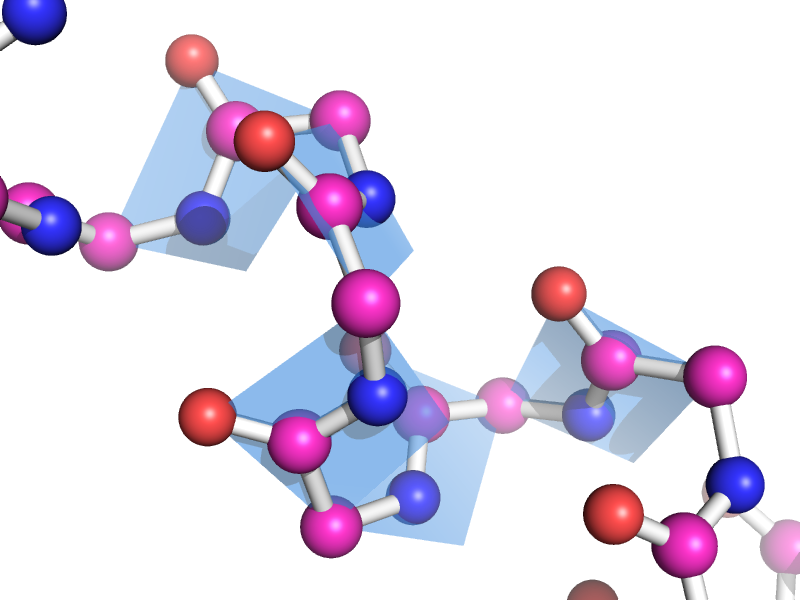
This script will draw a CGO plane between the backbone atoms of two neighboring residues. This is to show the planarity of the atoms. The image style this is meant to represent can be found many places, like "Introduction to Protein Structure" by Branden and Tooze (2nd ed. pp. 8).
Examples
# download the source and save as bbPlane.py
run bbPlane.py
fetch 1cll
# make planes for residues 4-9
bbPlane i. 4-10
The Source
#
# -- bbPlane.py - draws a CGO plane across the backbone atoms of
# neighboring amino acids
#
# Author: Jason Vertrees, 06/2010
# Modified by Thomas Holder, 06/2010
# Modified by Blaine Bell, 08/2011
# Copyright (C) Schrodinger
# Open Source License: MIT
#
from pymol.cgo import * # get constants
from pymol import cmd, stored
from chempy import cpv
def bbPlane(objSel='(all)', color='white', transp=0.0, state=1, name=None, quiet=1):
"""
DESCRIPTION
Draws a plane across the backbone for a selection
ARGUMENTS
objSel = string: protein object or selection {default: (all)}
color = string: color name or number {default: white}
transp = float: transparency component (0.0--1.0) {default: 0.0}
state = integer: object state, 0 for all states {default: 1}
NOTES
You need to pass in an object or selection with at least two
amino acids. The plane spans CA_i, O_i, N-H_(i+1), and CA_(i+1)
"""
# format input
transp = float(transp)
state, quiet = int(state), int(quiet)
if name is None:
name = cmd.get_unused_name("backbonePlane")
if state < 0:
state = cmd.get_state()
elif state == 0:
for state in range(1, cmd.count_states(objSel)+1):
bbPlane(objSel, color, transp, state, name, quiet)
return
stored.AAs = []
coords = dict()
# need hydrogens on peptide nitrogen
cmd.h_add('(%s) and n. N' % objSel)
# get the list of residue ids
for obj in cmd.get_object_list(objSel):
sel = obj + " and (" + objSel + ")"
for a in cmd.get_model(sel + " and n. CA", state).atom:
key = '/%s/%s/%s/%s' % (obj,a.segi,a.chain,a.resi)
stored.AAs.append(key)
coords[key] = [a.coord,None,None]
for a in cmd.get_model(sel + " and n. O", state).atom:
key = '/%s/%s/%s/%s' % (obj,a.segi,a.chain,a.resi)
if key in coords:
coords[key][1] = a.coord
for a in cmd.get_model(sel + " and ((n. N extend 1 and e. H) or (r. PRO and n. CD))", state).atom:
key = '/%s/%s/%s/%s' % (obj,a.segi,a.chain,a.resi)
if key in coords:
coords[key][2] = a.coord
# need at least two amino acids
if len(stored.AAs) <= 1:
print "ERROR: Please provide at least two amino acids, the alpha-carbon on the 2nd is needed."
return
# prepare the cgo
obj = [
BEGIN, TRIANGLES,
COLOR,
]
obj.extend(cmd.get_color_tuple(color))
for res in range(0, len(stored.AAs)-1):
curIdx, nextIdx = str(stored.AAs[res]), str(stored.AAs[res+1])
# populate the position array
pos = [coords[curIdx][0], coords[curIdx][1], coords[nextIdx][2], coords[nextIdx][0]]
# if the data are incomplete for any residues, ignore
if None in pos:
print 'peptide bond %s -> %s incomplete' % (curIdx, nextIdx)
continue
if cpv.distance(pos[0], pos[3]) > 4.0:
print '%s and %s not adjacent' % (curIdx, nextIdx)
continue
# need to order vertices to generate correct triangles for plane
# modified/added by B.Bell 8/18/2011
# modified by Thomas Holder 2012
if cpv.dot_product(cpv.sub(pos[0], pos[1]), cpv.sub(pos[2], pos[3])) < 0:
vorder = [0,1,2,2,3,0]
else:
vorder = [0,1,2,3,2,1]
# fill in the vertex data for the triangles;
for i in vorder:
obj.append(VERTEX)
obj.extend(pos[i])
# finish the CGO
obj.append(END)
# update the UI
cmd.load_cgo(obj, name, state)
cmd.set("cgo_transparency", transp, name)
cmd.extend("bbPlane", bbPlane)