|
|
Did you know...
With cartoon_gap_cutoff > 0, if there are missing residues along the protein backbone (e.g. missing loops), PyMOL will create a dashed cartoon loop segment if the gap (in number of residues) is shorter than the cutoff.
New in PyMOL 1.8.2
Example
set cartoon_gap_cutoff, 10
fetch 2xwu, async=0
as cartoon
orient B//152-156/
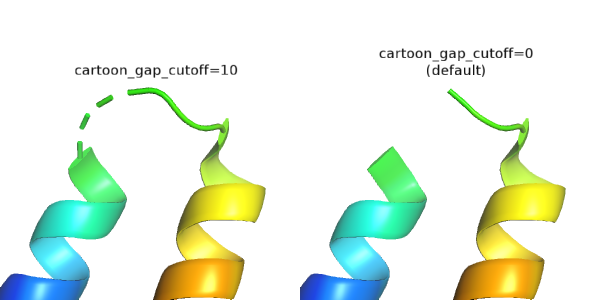
Number of dashes
The number of dashes is directly affected by the cartoon_sampling setting:
set cartoon_sampling, 20
See Also
|
|
|
 A Random PyMOL-generated Cover. See Covers.
|

