|
|
Did you know...
Overview
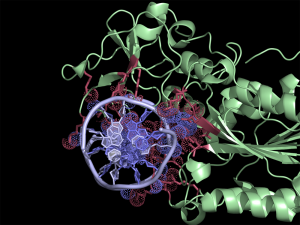 Interface residues (at cutoff <4A) in the 2c7r.pdb were found using NCONT, but similar results can be obtained using this script and CONTACT output. Usage of ccp4_contact script in PyMOL allows easy selection of residues and atoms listed in the output file. Interacting protein and DNA residues are colored in red and slate, respectively. Atoms in contact are shown in dots. The script selects residues and atoms from the list of the contacts found by CONTACT from CCP4 Program Suite (CONTACT analyses contacts between subsets of atoms in a PDB file).
First, we run CONTACT on our pdb file to find interface residues. Then by using the ccp4_contact script in PyMOL we separately select residues and atoms listed in the output file. This generates two selections (atoms and residues) for each interacting chain, allowing quick ..→
|
|
|
 A Random PyMOL-generated Cover. See Covers.
|

