Difference between revisions of "Visualizing a computed structure - a commented example"
Jump to navigation
Jump to search
Qqqqqqqqq9 (talk | contribs) |
|||
| Line 1: | Line 1: | ||
| − | + | # Obtain the [http://pubs.acs.org/cgi-bin/abstract.cgi/jacsat/2000/122/i37/abs/ja991878x.html data], the following structure was published in [http://pubs.acs.org/journals/jacsat/index.html J. Am. Chem. Soc.] in the Supporting Information | |
| − | in the Supporting Information | + | # Save [[Media:File.xyz.tar|File.xyz.tar]] and untar it. |
| − | + | # Convert the file | |
| − | + | ## convert it to pdb with [http://openbabel.org Openbabel] <source lang="python"> babel file.xyz file.pdb</source> | |
| − | + | ## Open it with [http://avogadro.openmolecules.net avogadro] and save it as pdb. Avogadro allows you to adjust the coordinate system, which saves some work if you want to visualize several similar structures. | |
| − | + | # Save the following pymolscript [[Media:script.pml.tar|Script.pml.tar]] and untar it to script.pml. Open it with an editor and adjust the Path_To_The_PDB-File. Open pymol and run the script with "@PATH_Of_The_Script/script.pml". | |
| − | + | # You will get the following image, you can save it with "png filename.png" [[Image:File.png]] | |
| − | babel file.xyz file.pdb | + | # Use the script to modify your own pdb-file. |
| − | |||
| − | |||
| − | |||
| − | Open it with | ||
| − | Avogadro allows you to adjust the coordinate | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | [[Image:File.png]] | ||
| − | |||
| − | |||
| − | |||
[[Category:Tutorials|Visualizing a computed structure - a commented example]] | [[Category:Tutorials|Visualizing a computed structure - a commented example]] | ||
Revision as of 09:01, 12 September 2008
- Obtain the data, the following structure was published in J. Am. Chem. Soc. in the Supporting Information
- Save File.xyz.tar and untar it.
- Convert the file
- Save the following pymolscript Script.pml.tar and untar it to script.pml. Open it with an editor and adjust the Path_To_The_PDB-File. Open pymol and run the script with "@PATH_Of_The_Script/script.pml".
- You will get the following image, you can save it with "png filename.png"
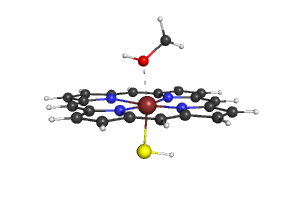
- Use the script to modify your own pdb-file.