Spectrum
The printable version is no longer supported and may have rendering errors. Please update your browser bookmarks and please use the default browser print function instead.
Overview
Spectrum colors atoms with a spectrum of colors based on an atomic property.
Usage
spectrum [expression [, palette [, selection [, minimum [, maximum [, byres ]]]]]]
expression
- count, b, q, or pc: respectively, atom count, temperature factor, occupancy, or partial charge {default: count}
palette
- string: palette name {default: rainbow}
selection
- string: atoms to color {default: (all)}
minimum
- float: {default: None (automatic)}
maximum
- float: {default: None (automatic)}
byres
- integer: controls whether coloring is applied per-residue {default: 0}
Notes
Minimum and maximum are determined automatically unless both arguments are provided and minimum < maximum.
Available palettes include:
- blue_green
- blue_magenta
- blue_red
- blue_white_green
- blue_white_magenta
- blue_white_red
- blue_white_yellow
- blue_yellow
- cbmr
- cyan_magenta
- cyan_red
- cyan_white_magenta
- cyan_white_red
- cyan_white_yellow
- cyan_yellow
- gcbmry
- green_blue
- green_magenta
- green_red
- green_white_blue
- green_white_magenta
- green_white_red
- green_white_yellow
- green_yellow
- green_yellow_red
- magenta_blue
- magenta_cyan
- magenta_green
- magenta_white_blue
- magenta_white_cyan
- magenta_white_green
- magenta_white_yellow
- magenta_yellow
- rainbow
- rainbow2
- rainbow2_rev
- rainbow_cycle
- rainbow_cycle_rev
- rainbow_rev red_blue
- red_cyan red_green
- red_white_blue
- red_white_cyan
- red_white_green
- red_white_yellow
- red_yellow
- red_yellow_green
- rmbc
- yellow_blue
- yellow_cyan
- yellow_cyan_white
- yellow_green
- yellow_magenta
- yellow_red
- yellow_white_blue
- yellow_white_green
- yellow_white_magenta
- yellow_white_red
- yrmbcg
Examples
Simple
spectrum b, blue_red, minimum=10, maximum=50
spectrum count, rainbow_rev, chain A, byres=1
Intermediate
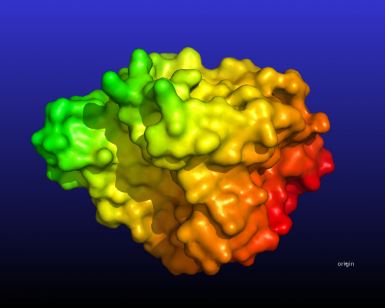
The following script will create this image:
# color atoms based on their distance from a point
# returns the length of the distance between atom A and atom B
diff_len = lambda x,y : math.sqrt((x[0]-y[0])*(x[0]-y[0]) + (x[1]-y[1])*(x[1]-y[1]) + (x[2]-y[2])*(x[2]-y[2]))
# fetch 1hug from the PDB
fetch 1hug, async=0
# show it as surface
as surface
# create the pseudoatom at the origin
pseudoatom pOrig, pos=(0,0,0), label=origin
# these are special PyMOL variables that will hold # the coordinates of
# the atoms and the pseudoatom
stored.origCoord = []
stored.distCoord = []
# copy the coordinates into those special variables
iterate_state 1, pOrig, stored.origCoord.append((x,y,z))
iterate_state 1, 1hug, stored.distCoord.append((x,y,z))
# extend origCoord to be the same length as the other
stored.origCoord *= len(stored.distCoord)
# calculate the distances
newB = map(lambda x,y: diff_len(x,y), stored.distCoord, stored.origCoord)
# put them into the b-factor of the protein
alter 1hug, b=newB.pop(0)
# color by rainbow_rev or any other
# palette listed in "help spectrum"
spectrum b, rainbow_rev, 1hug