Difference between revisions of "PyANM"
| Line 24: | Line 24: | ||
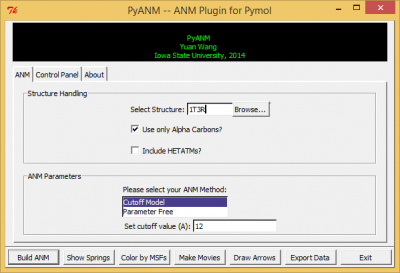
After a successful installation, go to Pymol-->Plugin and you will see PyANM, click on it and you should see PyANM's interface just like in figure 1. | After a successful installation, go to Pymol-->Plugin and you will see PyANM, click on it and you should see PyANM's interface just like in figure 1. | ||
| + | |||
| + | = Usage = | ||
| + | Now that you have PyANM installed, let's build one ANM together! | ||
| + | |||
| + | === Structure Handling === | ||
| + | *Type the name of your protein structure under 'Select Structure', this could either be a valid 4-letter PDB (Protein Data Bank) ID or the name of a object you have already loaded into Pymol. | ||
| + | *You could also choose to upload a local file from your system using the 'Browse' button. | ||
| + | *If the check box 'Use only Alpha Carbons' is checked, even if you choose an all-atom structure, PyANM will coarse-grain the structure for you. Otherwise PyANM will build an all-atom ANM if an all-atom structure is used. | ||
| + | *If the check box 'Include HETATMs' is checked, PyANM will include atoms in PDB that starts with HETATM as well. | ||
| + | |||
| + | === ANM Parameters === | ||
| + | *So far there are two ANMs available in PyANM : the cutoff model as described [http://www.cell.com/biophysj/abstract/S0006-3495%2801%2976033-X here] and the Parameter Free Model described [http://www.pnas.org/content/106/30/12347.short here], where spring constants are adjusted based on distance between two residues. | ||
| + | *If you choose to use <b>Cutoff Model</b>, you will be able to choose your desired cutoff value. The default value is 12 angstroms, personally I would recommend using any value between 7 angstroms to 15 anstroms. | ||
| + | *If you choose to use <b>Parameter Free Model</b>, you will be able to choose the power t for calculating spring constants for all residue pairs, where spring constant k for residue i and j is calculated as 1/distance(i,j) to the power of t. The default value for power t is 6. | ||
Revision as of 18:21, 19 October 2014
| Type | PyMOL Plugin |
|---|---|
| Download | plugins/pyanm.py |
| Author(s) | Yuan Wang |
| License | - |
| This code has been put under version control in the project Pymol-script-repo | |
Introduction
Elastic Network Models (ENM) have been successful in reproducing fluctuations for proteins of native conformations. ENM is a coarse-grained method for modeling protein dynamics, meaning in generally in ENM each residue is represented by one bead in its Alpha Carbon position. These beads are then connected by elastic springs if the distance between two beads fall under a cutoff value (usually within the range of 7 to 15 angstroms). For more details please read here.
PyANM is developed as a cross-platform Pymol Plugin to allow its users to build and visualize Anisotropic Network Models, a member of the ENM family. This plugin allows its users to draw arrows or make movies based on calculated mode motions, to draw all springs used to build the ANM within the protein, to color the protein based on its Mean Square Fluctuations (MSF) calculated from ANM and to export data from ANM for future processing. PyANM also allows its users to change the colors and scales (sizes) for movies or arrows generated by PyANM for better visualization.
Dependencies
- PyANM requires Numerical Python (NumPy) to do all the matrix calculations. If you wish to use PyANM but don't have NumPy on your machine, please follow the download and build instructions here. If you will be using PyANM on a Windows machine, installing a Python distribution such as Anaconda might be the easier. After installing NumPy, type the following line inside Pymol and if you don't see any error messages, you now have NumPy ready for Pymol!
import numpy
- Having SciPy might improve the performance of PyANM but it is not required.
- PyANM has only been tested for Pymol 1.1 and higher, using an earlier version on Pymol might cause problems.
Installation
Download pyanm.py from this page, then go to Pymol-->Plugin-->Manage Plugins-->Install..., select the file you have just downloaded and then restart Pymol. Things might be different depending on your Pymol version and your local system, but this should be fairly similar.
After a successful installation, go to Pymol-->Plugin and you will see PyANM, click on it and you should see PyANM's interface just like in figure 1.
Usage
Now that you have PyANM installed, let's build one ANM together!
Structure Handling
- Type the name of your protein structure under 'Select Structure', this could either be a valid 4-letter PDB (Protein Data Bank) ID or the name of a object you have already loaded into Pymol.
- You could also choose to upload a local file from your system using the 'Browse' button.
- If the check box 'Use only Alpha Carbons' is checked, even if you choose an all-atom structure, PyANM will coarse-grain the structure for you. Otherwise PyANM will build an all-atom ANM if an all-atom structure is used.
- If the check box 'Include HETATMs' is checked, PyANM will include atoms in PDB that starts with HETATM as well.
ANM Parameters
- So far there are two ANMs available in PyANM : the cutoff model as described here and the Parameter Free Model described here, where spring constants are adjusted based on distance between two residues.
- If you choose to use Cutoff Model, you will be able to choose your desired cutoff value. The default value is 12 angstroms, personally I would recommend using any value between 7 angstroms to 15 anstroms.
- If you choose to use Parameter Free Model, you will be able to choose the power t for calculating spring constants for all residue pairs, where spring constant k for residue i and j is calculated as 1/distance(i,j) to the power of t. The default value for power t is 6.