Pseudoatom
pseudoatom creates a molecular object with a pseudoatom or adds a pseudoatom to a molecular object if the specified object already exists. Default position is in the middle of the viewing window.
USAGE
pseudoatom object [, selection [, name [, resn [, resi [, chain
[, segi [, elem [, vdw [, hetatm [, b [, q [, color [, label
[, pos [, state [, mode [, quiet ]]]]]]]]]]]]]]]]]
You can set the following: selection, name, resn, resi, chain, segi, element, vdw, hetatm, b-factor, charge, color, label, pos (location in space), state, mode, and quiet.
# create the pseudoatom
pseudoatom tmpPoint
ObjMol: created tmpPoint/PSDO/P/PSD`1/PS1
# show it as a sphere.
show spheres, tmpPoint
# create another, with more options.
pseudoatom tmpPoint2, resi=40, chain=ZZ, b=40, color=tv_blue, pos=[-10, 0, 10]
ObjMol: created tmpPoint2/PSDO/ZZ/PSD`40/PS1
EXAMPLES FOR USE
pseudoatom can be used for a wide variety of tasks where on must place an atom or a label in 3D space, e.g. as a placeholder for distance measurement or distance specifications.
# A pseudoatom as a placeholder for selections according to distance:
load $TUT/1hpv.pdb
pseudoatom tmp, pos=[10.0, 17.0, -3.0]
show sticks, tmp expand 6
delete tmp
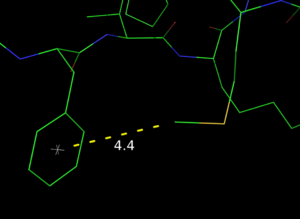
# A pseudoatom as placeholder for distance measurement:
# position it at the center of an aromatic ring. Then
# calc the distance from another atom to the pseudoatom.
load $TUT/1hpv.pdb
pseudoatom pi_cent,b/53/cg+cz
dist pi_cent////ps1, b/met`46/ce
You can use a pseudoatom to make a label for a scene title. Move the protein to the bottom of the window before the pseudoatom is created. Or move the label after creating it (Shift + Middle mouse button in editing mode).
fetch 1rq5
pseudoatom forLabel
label forLabel, "This Protein is a Carbohydrate Active Enzyme"
set label_color, black
# png ray=1
References
PyMOL Mailing List