Difference between revisions of "Main Page"
(PyMOL 3.0 release) |
|||
| (65 intermediate revisions by 14 users not shown) | |||
| Line 1: | Line 1: | ||
__NOTOC__ | __NOTOC__ | ||
| − | {| align="center" style="padding-bottom: | + | {| align="center" width="100%" style="background: #B22222; margin-bottom: 4em; border-bottom: 1px solid #B22222; border-left: 1px solid #B22222; border-right: 1px solid #B22222;" |
| − | |+ style="font-size:210%; font-weight: bold; color:# | + | |+ style="font-size: 1.0em; font-weight: normal; color: #FFFFFF; text-align:right; background: #B22222; padding-top:0.5em; padding-bottom: 0.25em; padding-right: 0.60em; border-top: 2px solid #B22222; border-bottom: 1px solid #fff;" |hosted by [[File:SBGridlogo2.jpg|140 px|link=https://sbgrid.org/]] |
| − | |- style="text-align:center; font-weight: | + | |} |
| + | {| align="center" style="padding-bottom: 3em;" | ||
| + | |+ style="font-size:210%; font-weight: bold; color:#000000; text-align:center; padding: 5px; margin-bottom: 4px;" | Welcome to the PyMOL Wiki! | ||
| + | |- style="text-align:center; font-weight: normal; color: #000000; font-size: 120%; font-family: sans-serif;" | ||
| The community-run support site for the [http://pymol.org PyMOL] molecular viewer. | | The community-run support site for the [http://pymol.org PyMOL] molecular viewer. | ||
| − | |- style="text-align:center; font-weight: | + | |- style="text-align:center; font-weight:normal; color: #000000; font-size: 120%; font-family: sans-serif;" |
| − | | | + | | To request a new account, email SBGrid at: accounts (@) sbgrid dot org |
| − | |- style="text-align:center; font-weight:bold; color: # | + | |- style="text-align:center; font-weight:bold; color: #000000; font-size: 120%; font-family: sans-serif;" |
|} | |} | ||
| − | {| align="center" width="45%" style="background: # | + | |
| − | |+ style="font-size: 1.4em; font-weight: bold; color: # | + | {| align="center" width="45%" style="background: #FFFFFF; margin-bottom: 4em; border-bottom: 1px solid #AFB29E; border-left: 1px solid #AFB29E; border-right: 1px solid #AFB29E;" |
| + | |+ style="font-size: 1.4em; font-weight: bold; color: #FFFFFF; text-align:center; background: #000000; padding-top:0.5em; padding-bottom: 0.25em; border-top: 2px solid #000000; border-bottom: 1px solid #fff;" |Quick Links | ||
|- | |- | ||
| − | | style="font-size: 1.1em; color # | + | | style="font-size: 1.1em; font-weight: normal; color #48A2B4; padding: 0.5em 1em 0.5em 3em;"|'''[[:Category:Tutorials|Tutorials]]''' || '''[[TOPTOC|Table of Contents]]''' || '''[[:Category:Commands|Commands]]''' |
|- | |- | ||
| − | | style="font-size: 1.1em; color # | + | | style="font-size: 1.1em; font-weight: normal; color #48A2B4; padding: 0.5em 1em 0.5em 3em;"|'''[[:Category:Script_Library|Script Library]]''' || '''[[:Category:Plugins|Plugins]]''' || '''[[:Category:FAQ|FAQ]]''' |
|- | |- | ||
| − | | style="font-size: 1.1em; color # | + | | style="font-size: 1.1em; font-weight: normal; color #48A2B4; padding: 0.5em 1em 0.5em 3em;"|'''[[Gallery]]''' | '''[[Covers]]''' |
||'''[[CheatSheet|PyMOL Cheat Sheet]]''' (''[[Media:PymolRef.pdf|PDF]]'') | ||'''[[CheatSheet|PyMOL Cheat Sheet]]''' (''[[Media:PymolRef.pdf|PDF]]'') | ||
| − | ||'''[[ | + | ||'''[[PyMOL_mailing_list|Getting Help]]''' |
|} | |} | ||
| Line 25: | Line 29: | ||
|+ style="font-size: 1.4em; font-weight: bold; text-align:left; border-bottom: 2px solid #6678b1;" | News & Updates | |+ style="font-size: 1.4em; font-weight: bold; text-align:left; border-bottom: 2px solid #6678b1;" | News & Updates | ||
|- | |- | ||
| − | ! | + | ! Official Release |
| − | | | + | | [https://pymol.org PyMOL v3.0 has been released] on March 12, 2024. |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
|- | |- | ||
! New Plugin | ! New Plugin | ||
| − | | [[ | + | | [[CavitOmiX|CavitOmiX]] calculate [https://innophore.com Catalophore™ cavities], predict protein structures with [https://www.nvidia.com/en-us/gpu-cloud/bionemo OpenFold by NVIDIA-BioNeMo], [https://ai.facebook.com/blog/protein-folding-esmfold-metagenomics/ ESMFold] and retrieve [https://www.deepmind.com/research/highlighted-research/alphafold Alphafold] models |
|- | |- | ||
| − | ! | + | ! Official Release |
| − | | [ | + | | [https://pymol.org PyMOL v2.5 has been released] on May 10, 2021. |
|- | |- | ||
| − | ! | + | ! Python 3 |
| − | | [[ | + | | New [[2to3|Python 3 compatibility guide]] for scripts and plugins |
|- | |- | ||
| − | ! | + | ! POSF |
| − | | [ | + | | [https://pymol.org/fellowship New PyMOL fellowship announced for 2022-2023] |
|- | |- | ||
| − | ! | + | ! Tutorial |
| − | | [[ | + | | [[Plugins Tutorial]] updated for PyQt5 |
|- | |- | ||
! New Plugin | ! New Plugin | ||
| − | | [[ | + | | [[PICv|PICv]] is a new plugin for clustering protein-protein interactions and visualization with available data from PDBe |
|- | |- | ||
| − | ! | + | ! Selection keywords |
| − | | [[ | + | | New [[Selection Algebra|polymer.protein and polymer.nucleic]] selection keywords. Thanks everyone who participated in the [https://goo.gl/forms/r0Ck03VTytZQxN4A2 poll]! |
|- | |- | ||
| − | ! | + | ! Plugin Update |
| − | | [[ | + | | [[MOLE 2.0: advanced approach for analysis of biomacromolecular channels|MOLE 2.5]] is an updated version of channel analysis software in PyMOL |
| − | |||
| − | |||
| − | |||
|- | |- | ||
! New Script | ! New Script | ||
| − | | [[ | + | | [[dssr_block]] is a wrapper for DSSR (3dna) and creates block-shaped nucleic acid cartoons |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
|- | |- | ||
! Older News | ! Older News | ||
| Line 96: | Line 67: | ||
|- | |- | ||
|<div class="didyouknow" > | |<div class="didyouknow" > | ||
| − | < | + | <DynamicPageList> |
| − | + | randomcount=1 | |
category=Commands|Plugins|Script_Library|Settings | category=Commands|Plugins|Script_Library|Settings | ||
includepage=* | includepage=* | ||
| − | includemaxlength= | + | includemaxlength=1050 |
escapelinks=false | escapelinks=false | ||
| + | allowcachedresults=false | ||
resultsheader=__NOTOC__ __NOEDITSECTION__ | resultsheader=__NOTOC__ __NOEDITSECTION__ | ||
| − | |||
| − | |||
| − | |||
listseparators=,<h3>[[%PAGE%]]</h3>,,\n | listseparators=,<h3>[[%PAGE%]]</h3>,,\n | ||
| − | </ | + | </DynamicPageList> |
</div> | </div> | ||
<div style="clear: both;"></div> | <div style="clear: both;"></div> | ||
| Line 113: | Line 82: | ||
| | | | ||
|style="vertical-align: top; width: 18%"| | |style="vertical-align: top; width: 18%"| | ||
| − | < | + | <DynamicPageList> |
imagecontainer=Covers | imagecontainer=Covers | ||
randomcount=1 | randomcount=1 | ||
| Line 120: | Line 89: | ||
listseparators=[[,%PAGE%,|thumb|185px|A Random PyMOL-generated Cover. See [[Covers]].]],\n | listseparators=[[,%PAGE%,|thumb|185px|A Random PyMOL-generated Cover. See [[Covers]].]],\n | ||
ordermethod=none | ordermethod=none | ||
| − | </ | + | allowcachedresults=false |
| + | </DynamicPageList> | ||
|} | |} | ||
Latest revision as of 12:54, 12 March 2024
| The community-run support site for the PyMOL molecular viewer. |
| To request a new account, email SBGrid at: accounts (@) sbgrid dot org |
| Tutorials | Table of Contents | Commands |
| Script Library | Plugins | FAQ |
| Gallery | Covers | PyMOL Cheat Sheet (PDF) | Getting Help |
|
|
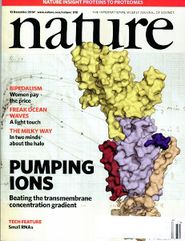 A Random PyMOL-generated Cover. See Covers.
|