Gallery
Jump to navigation
Jump to search
The printable version is no longer supported and may have rendering errors. Please update your browser bookmarks and please use the default browser print function instead.
| Cool PyMOL-generated Images and their Scripts. Add Your Own |
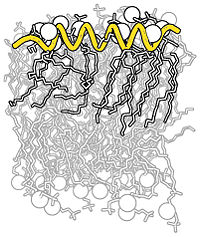
| Complex B&W outline representation | What To Type | |||||
|
# first load lipid model
load lipids.pdb;
# hide the initially loaded representation
hide all;
# set background color to white
bg_color white;
# show lipid model as sticks
show sticks, lipids;
# color the lipids model by element CHNOS #2 (carbon green)
util.cbag lipids;
# select all hydrogens and remove them from the model
select hideme, hydro;
hide everything, hideme;
delete hideme;
# create phosphate spheres
create phos, elem p;
hide everything, phos;
show spheres, phos;
# load helix model
load helix.pdb;
# hide the initially loaded representation
hide everything, helix;
# make the helical struct into a cartoon form
show cartoon, helix;
# style the cartoon form
cartoon putty;
# reposition the helix among the lipids using
# the 3-Button Editing Mouse Mode
# basically
# Shift+Left Mouse to rotate the helix
# Shift+Middle Mouse to move the helix
# also, you may want to make liberal use of the
# get_view and set_view commands.
#
# When you have the scene set like you want,
# continue with...
# move the model to find the view you want,
# and use get_view to get the coordinate description
get_view;
# set ray_trace_mode to black and white outline
set ray_trace_mode, 2;
|
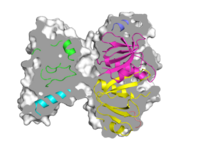
| Grid Mode | What To Type | |||||
|
fetch 1cll 1sra 1ggz 5pnt 1rlw 1cdy;
set grid_mode
Hint: You may wish to execute the 'reset' command on the command line after running the above commands to get full molecules in view of window and centered in a more useable manner. |
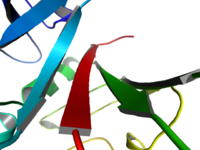
| Cool Perspective | What To Type | |||||
|
load prot.pdb;
zoom i. 46-49 and n. CA
set field_of_view, 60
ray
|
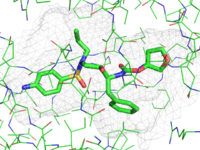
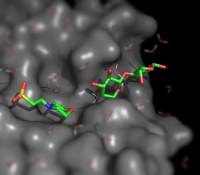
| Representing a binding pocket | What To Type | |||||
|
load $TUT/1hpv.pdb, tmp
extract lig, organic
extract prot, polymer
delete tmp
set surface_carve_cutoff, 4.5
set surface_carve_selection, lig
set surface_carve_normal_cutoff, -0.1
show surface, prot within 8 of lig
set two_sided_lighting
set transparency, 0.5
show sticks, lig
orient lig
set surface_color, white
set surface_type, 2 # mesh
unset ray_shadows
|
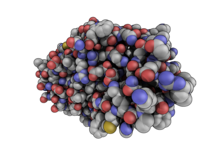
| QuteMol Like | What To Type | |||||
|
load $TUT/1hpv.pdb
set_color oxygen, [1.0,0.4,0.4]
set_color nitrogen, [0.5,0.5,1.0]
remove solvent
as spheres
util.cbaw
bg white
set light_count,8
set spec_count,1
set shininess, 10
set specular, 0.25
set ambient,0
set direct,0
set reflect,1.5
set ray_shadow_decay_factor, 0.1
set ray_shadow_decay_range, 2
unset depth_cue
# for added coolness
# set field_of_view, 60
ray
|
| Simulating Tilt-shift | What To Type | |||||
|
fetch 1wld
as surface, poly
as sticks, org
h_add solvent
color grey, poly
orient org
png img.png
# now, go into Photoshop or the GIMP and apply a Gaussian or
# Focus blur to the top and bottom portions of the image
|
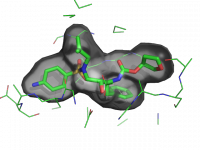
| Ray-normal-based transparency | What To Type | |||||
|
# grey surface
set surface_color, grey
# cavity mode
set surface_mode, 3
# layered transparency mode
set transparency_mode, 1
# surface transparency
set transparency, 0.5
# oblique and contrast define the
# look of the surface transparency:
# if the normal vector is
set ray_transparency_oblique
set ray_transparency_oblique_power, 8
set ray_transparency_contrast, 7
# fetch a protein, with a
# small molecule in a nice
# hidden pocket
fetch 1hpv, async=0
hide
# show the small molecule as surface
show surface, org
# arrange the view
orient org
# zoom back a little
zoom org, 1
# show the small molecule inside as sticks
show sticks, org
# show some nearby sidechains
show lines, poly within 5 of org
# enable frame caching for playback
set cache_frames, 1
set ray_trace_frames, 1
mset 1x120
movie.roll 1, 120, 1, x
mplay
# now sit back and wait 5 minutes...
|
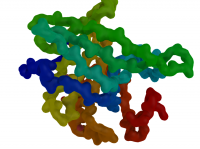
| Blobby Main Chain | What To Type | |||||
|
fetch 3uex, struct, async=0
remove solvent
# set the B-factors nice and high for smoothness
alter all, b=10
alter all, q=1
# 3.5 A map resolution
set gaussian_resolution, 3.5
# new gaussian map w/resolution=0.5 Ang
# on just the main chain
map_new map, gaussian, 0.5, n. C+O+N+CA, 5
# create a surface from the map
isosurface surf, map, 3.0
# color the protein by number
spectrum count, rainbow, struct
# now color the map based on the underlying protein
cmd.ramp_new("ramp", "struct", [0,10,10], [-1, -1, 0])
# set the surface color
cmd.set("surface_color", "ramp", "surf")
# hide the ramp and lines
disable ramp
hide lines
bg grey
# soften out the image
set light_count,8
set spec_count,1
set shininess, 10
set specular, 0.075
set ambient,0
set direct,0
set reflect, 0.85
set ray_shadow_decay_factor, 0.1
set ray_shadow_decay_range, 4
unset depth_cue
# ray trace the image
orient
ray
|
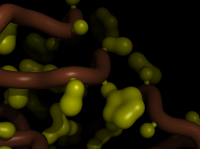
| Blobby Side Chains | What To Type | |||||
|
# fetch a protein
fetch 1rx1, async=0
# setup the cartoon tubes
as cartoon
cartoon tube
set cartoon_tube_radius, 0.7
set cartoon_color, brown
set cartoon_side_chain_helper, on
show sticks, poly
color yellow
# README
# stop here, or try this for "sloppy sticks"
# beefy video card required!
select rep sticks
select sele and not n. C+O+N+CA
# set the B-factors nice and high for smoothness
alter all, b=10
alter all, q=1
# 2.5 A map resolution
set gaussian_resolution, 2.5
# 0.2 A sampling; lower=smoother
map_new map, gaussian, 0.2, sele, 5
# create a surface from the map
isosurface surf, map, 5.0
color yellow, surf
hide sticks
# reconnect the main chain to the blobs
show sticks, n. CA+CB
|
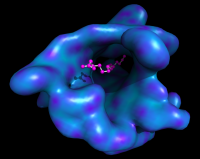
| Smooth surface with ligand | What To Type | |||||
|
fetch 3uex, struct, async=0
remove solvent
# set the B-factors nice and high for smoothness
alter all, b=10
alter all, q=1
# 3.5 A map resolution
set gaussian_resolution, 7.6
# new gaussian map w/resolution=0.5 Ang
# on just the main chain
map_new map, gaussian, 1, n. C+O+N+CA, 5
# create a surface from the map
isosurface surf, map, 1.5
# color the protein by number
spectrum count, rainbow, struct
# now color the map based on the b-factors of the
# underlying protein
cmd.ramp_new("ramp", "struct", [0,10,10], "rainbow")
# set the surface color
cmd.set("surface_color", "ramp", "surf")
# hide the ramp and lines
disable ramp
hide lines
show sticks, org
show spheres, org
color magenta, org
reinit
fetch 3uex, struct, async=0
remove solvent
# set the B-factors nice and high for smoothness
alter all, b=10
alter all, q=1
# 3.5 A map resolution
set gaussian_resolution, 7.6
# new gaussian map w/resolution=0.5 Ang
# on just the main chain
map_new map, gaussian, 1, n. C+O+N+CA, 5
# create a surface from the map
isosurface surf, map, 1.5
# color the protein by number
spectrum count, rainbow, struct
# now color the map based on the b-factors of the
# underlying protein
cmd.ramp_new("ramp", "struct", [0,10,10], "rainbow")
# set the surface color
cmd.set("surface_color", "ramp", "surf")
# hide the ramp and lines
disable ramp
hide lines
show sticks, org
show spheres, org
color magenta, org
set_bond stick_radius, 0.13, org
set sphere_scale, 0.26, org
set_bond stick_radius, 0.13, org
set_bond stick_color, white, org
set sphere_scale, 0.26, org
set_view (\
-0.877680123, 0.456324875, -0.146428943,\
0.149618521, -0.029365506, -0.988305628,\
-0.455291569, -0.889327347, -0.042500813,\
-0.000035629, 0.000030629, -37.112102509,\
-3.300258160, 6.586110592, 22.637466431,\
8.231912613, 65.999290466, -50.000000000 )
# ray trace the image
ray
|
| Complex Stylized Protein | What To Type | |||||
|
fetch 1eaz, async=0
extract oo, org
hide everything, solvent
set field_of_view, 50
preset.ball_and_stick("oo")
set_bond stick_color, 0xffff44, oo
set_bond stick_transparency, 0.35, oo
color grey, oo and e. C
set valence, 1, oo
ramp_new pRamp, oo, selection=poly, range=[5,30], color=rainbow
set surface_color, pRamp, poly
show spheres, poly
color white, poly
color grey30, poly and e. C
set sphere_scale, 0.99, poly
set ray_transparency_contrast, 0.20
set ray_transparency_oblique, 1.0
set ray_transparency_oblique_power, 20
show surface, poly
set surface_quality, 2
set light_count, 5
set ambient_occlusion_mode, 1
set ambient_occlusion_scale, 50
set ambient, 0.40
set transparency, 0.50
disable pRamp
set spec_power, 1200
set spec_reflect, 0.20
set ray_opaque_background, 0
set ray_shadow, 0
ray
|
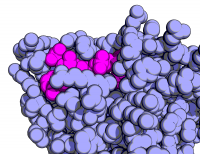
| Goodsell-like | What To Type | |||||
|
# fetch the protein
fetch 1rx1, async=0
# show it as blue/magenta spheres
as spheres
color lightblue, not org
color magenta, org
remove solvent
# set the view
orient all within 8 of org
# set the lights, ray tracing setttings
# to get the Goodsell-like rendering
unset specular
set ray_trace_gain, 0
set ray_trace_mode, 3
bg_color white
set ray_trace_color, black
unset depth_cue
ray
|
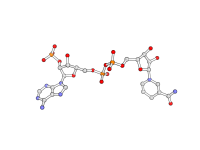
| Stylized Ball and Stick | What To Type | |||||
|
hide everything
show sticks
show spheres
set stick_radius, .07
set sphere_scale, .18
set sphere_scale, .13, elem H
set bg_rgb=[1, 1, 1]
set stick_quality, 50
set sphere_quality, 4
color gray85, elem C
color red, elem O
color slate, elem N
color gray98, elem H
set stick_color, black
set ray_trace_mode, 1
set ray_texture, 2
set antialias, 3
set ambient, 0.5
set spec_count, 5
set shininess, 50
set specular, 1
set reflect, .1
set dash_gap, 0
set dash_color, black
set dash_gap, .15
set dash_length, .05
set dash_round_ends, 0
set dash_radius, .05
python
preset.ball_and_stick("vis")
python end
fetch 1rx1, async=0
remove not org
orient org
ray
|
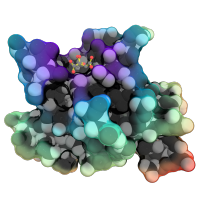
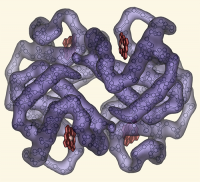
| Tribute to Irving Geis | What To Type | |||||
|
# I've always appreciated the simplicity of Irving Geis's designs.
# Here, I've reproduced one of his famous images of hemoglobin.
fetch 1buw
remove solvent
hide all
# Irving Geis shows only the carbonyl carbon atoms in his structure,
# so build a backbone made up of only those atoms
create bbC_A, n. C in /1buw/A/A
stored.bbC = []
iterate (bbC_A), stored.bbC.append(index)
for i in stored.bbC: cmd.bond("i. %s in bbC_A" % str(i), "i. %s in bbC_A" % str(i+1))
create bbC_B, n. C in /1buw/B/B
stored.bbC = []
iterate (bbC_B), stored.bbC.append(index)
for i in stored.bbC: cmd.bond("i. %s in bbC_B" % str(i), "i. %s in bbC_B" % str(i+1))
create bbC_C, n. C in /1buw/C/C
stored.bbC = []
iterate (bbC_C), stored.bbC.append(index)
for i in stored.bbC: cmd.bond("i. %s in bbC_C" % str(i), "i. %s in bbC_C" % str(i+1))
create bbC_D, n. C in /1buw/D/D
stored.bbC = []
iterate (bbC_D), stored.bbC.append(index)
for i in stored.bbC: cmd.bond("i. %s in bbC_D" % str(i), "i. %s in bbC_D" % str(i+1))
show sticks, /bbC_A or /bbC_B or /bbC_C or /bbC_D
set stick_radius, 0.2
show spheres, bbC_A or bbC_B or bbC_C or bbC_D
set sphere_scale, 0.4
color white, bbC_A or bbC_B or bbC_C or bbC_D
# Create isosurface maps to draw the backbone surface as a tube
set gaussian_b_floor, 40
set gaussian_resolution, 5
map_new mapA, gaussian, 1, bbC_A, 60
isosurface isoA, mapA, 15
map_new mapB, gaussian, 1, bbC_B, 60
isosurface isoB, mapB, 15
map_new mapC, gaussian, 1, bbC_C, 60
isosurface isoC, mapC, 15
map_new mapD, gaussian, 1, bbC_D, 60
isosurface isoD, mapD, 15
set transparency, 0.2
# Create a color gradient with a color ramp from a pseudoatom
set_color tubedark, [67, 45, 133]
set_color tubelight, [178, 177, 204]
pseudoatom pseud, pos=[142.982, 26.505, 86.321]
hide /pseud
ramp_new colorRamp, pseud, range=[0,100,142], color=[tubedark, tubedark, tubelight]
set surface_color, colorRamp, isoA
set surface_color, colorRamp, isoB
set surface_color, colorRamp, isoC
set surface_color, colorRamp, isoD
disable colorRamp
# Represent the heme rings as cartoons
create hemes, /1buw/E/A or /1buw/G/B or /1buw/I/C or /1buw/K/D
# Recreate a lost bond...
bond /hemes/K/D/HEM`147/FE, /hemes/K/D/HEM`147/NC
show_as cartoon, hemes
set cartoon_ring_finder, 4
set cartoon_ring_mode, 3
set cartoon_ring_width, 0.3
set cartoon_ring_transparency, 0
color ruby, hemes
# Represent the heme iron ions as spheres
create irons, e. Fe
show_as spheres, irons
set sphere_scale, 0.6, irons
color darksalmon, irons
# Hide ligands bound to hemes
hide //F or //H or //J or //L
# Hide atoms that stick out of the surface (mostly terminal atoms)
hide /bbC_C/C/C/141
hide /bbC_A/A/A/141
hide /bbC_D/D/D/144
hide /bbC_D/D/D/1
# Display settings
set_color bground, [252, 247, 229]
bg_color bground
set field_of_view, 5
set_view (\
0.568997085, 0.032208484, 0.821707368,\
0.172765702, 0.972248971, -0.157743633,\
-0.803984761, 0.231720015, 0.547644258,\
0.000000000, 0.000000000, -691.255310059,\
48.556941986, 45.649765015, 22.998313904,\
579.895507812, 802.615112305, 5.000000000 )
set ray_trace_mode, 1
set ray_shadow, 0
set light_count, 2
set light, [0, 0, -100]
set ambient, 0
set direct, 0.7
set specular, 1
set shininess, 5
set specular_intensity, 0.3
set reflect, 0.2
set reflect_power, 1
set depth_cue, 1
set fog_start, 0.45
set antialias, 3
ray 1280, 960
|