The printable version is no longer supported and may have rendering errors. Please update your browser bookmarks and please use the default browser print function instead.
Default settings
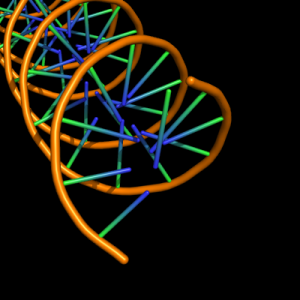
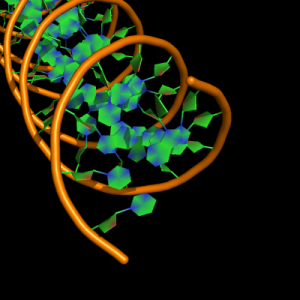
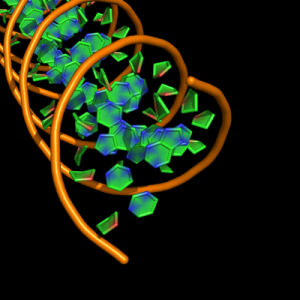
The defaults give a phosphate backbone with single sticks passing across the full width of the base plane.
set cartoon_nucleic_acid_mode, 0 # backbone follows phosphates; actually Pymol itself uses setting '4' as default
set cartoon_ladder_mode, 1 # sticks from backbone into nucleotide
set cartoon_ring_mode, 0 # no nucleotide rings
set cartoon_ring_finder, 1 # ribose and base rings (not displayed since ring mode 0)
Cartoon ring mode
Settings
set cartoon_ring_mode, value
| value |
effect
|
| 0 |
stick from backbone atom to N1 of purines and N3 of pyrimidines
|
| 1 |
simple plane for ribose and base rings covering area between ring bonds
|
| 2 |
simple plane for ribose and base rings covering area inside sticks (slightly smaller than mode 1)
|
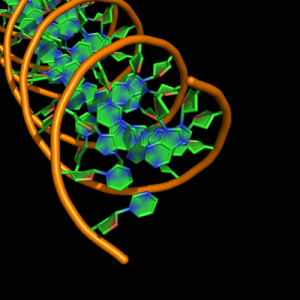
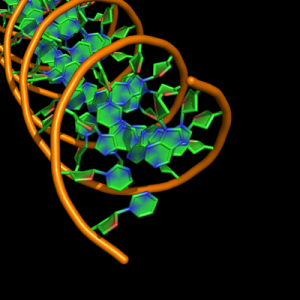
| 3 |
plane bounded by sticks for ribose and base rings
|
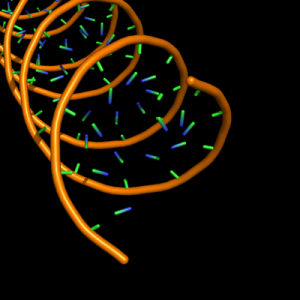
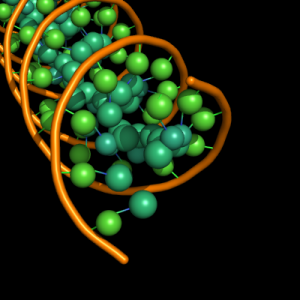
| 4 |
large sphere of ring diameter at centre of ribose and each base ring
|
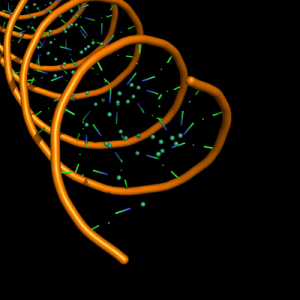
| 5 |
small sphere of 1/10 diameter at centre of ribose and each base ring
|
Examples
all with defaults: cartoon_ladder_mode,1 cartoon_nucleic_acid_mode,0 cartoon_ring_finder,1
Cartoon ring finder
Settings
set cartoon_ring_finder, value
| value |
effect
|
| 0 |
no rings or sticks joining them
|
| 1 |
both ribose and base ring
|
| 2 |
only base ring(s), stick connects directly from phosphate to ring
|
| 3 |
very similar to ring finder 1, slight effect on transparency = distinct behaviour?
|
| 4 |
very similar to ring finder 1: finds ribose and base of nucleotides, and aromatic side chains of proteins
|
| 5 |
sticks visible but rings invisible
|
Examples
all with: cartoon_ladder_mode,1 cartoon_ring_mode,3 cartoon_nucleic_acid_mode,0
Cartoon ladder mode
Settings
set cartoon_ladder_mode, value
| value |
effect
|
| 0 |
no sticks shown
|
| 1 |
sticks show
|
Examples
all with: cartoon_ring_mode,3 cartoon_nucleic_acid_mode,0 cartoon_ring_finder,1
Note that the visibility of the ladder sticks depends on ring mode >0, ring finder >0, nucleic acid mode = 0
Cartoon nucleic acid mode
Settings
set cartoon_nucleic_acid_mode, value
| value |
effect
|
| 0 |
smooth backbone passing through phosphorus atoms, backbone terminates at last phosphorus on either end of chain
|
| 1 |
smooth backbone passing through ribose C3' atoms, backbone terminates at last C3' on either end of chain
|
| 2 |
smooth backbone passing through phosphorus atoms, backbone terminates at last phosphorus on 5' end and O3' on 3' end (note takes O3' colour at terminus in default colouring)
|
| 3 |
appears same as mode 0?
|
| 4 |
appears same as mode 2? Seems to be what Pymol uses when it first opens nucleic acid containing file because any other settings change ends and colors.
|
Examples
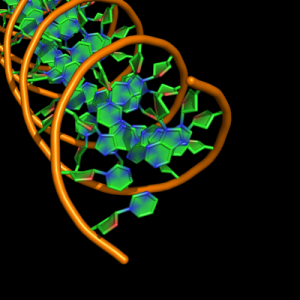 cartoon_nucleic_acid_mode,0 |
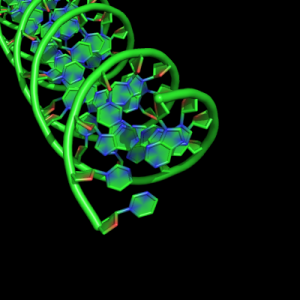 cartoon_nucleic_acid_mode,1 |
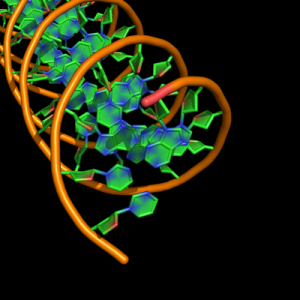 cartoon_nucleic_acid_mode,2 |
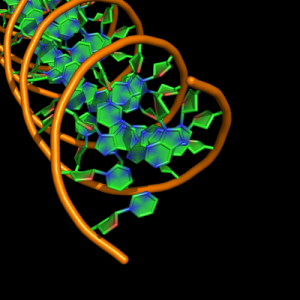 cartoon_nucleic_acid_mode,3 |
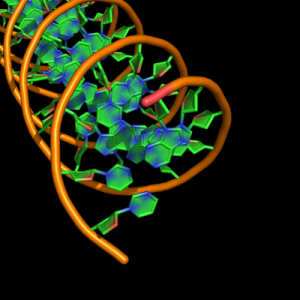 cartoon_nucleic_acid_mode,4 |
all with: cartoon_ladder_mode,0 cartoon_ring_mode,3 cartoon_ring_finder,1