Difference between revisions of "Examples of nucleic acid cartoons"
Jump to navigation
Jump to search
all with defaults: cartoon_ladder_mode,1 cartoon_nucleic_acid_mode,0 cartoon_ring_finder,1
(No difference)
| |
Revision as of 12:57, 9 December 2007
pymol version info
The various new nucleic acid display settings for PyMol 0.99 listed below were produced using version 0.99 beta 29 on Windows XP with an arbitrary B-form DNA molecule. Higher values could be set for each setting but appeared to yield the default representations.
default settings
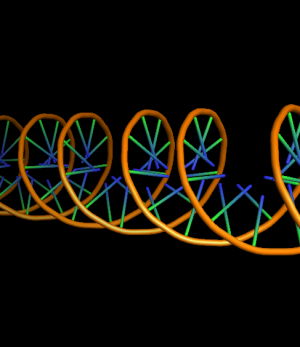
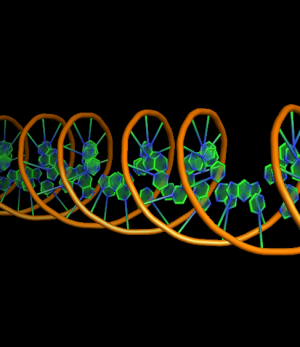
The defaults give a phosphate backbone with single sticks passing across the full width of the base plane.
set cartoon_nucleic_acid_mode, 1 # backbone follows phosphates
set cartoon_ladder_mode, 1 # sticks from backbone into nucleotide
set cartoon_ring_mode, 0 # no nucleotide rings
set cartoon_ring_finder, 1 # ribose and base rings (not displayed as ring mode 0)
cartoon ring modes
set cartoon_ring_mode, 0 # no nucleotide rings
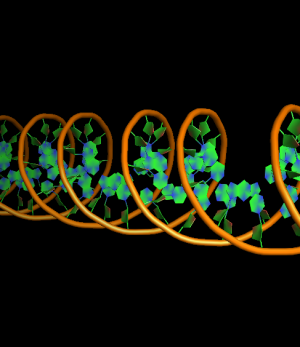
set cartoon_ring_mode, 1 # filled rings extending to outside edge of bonds
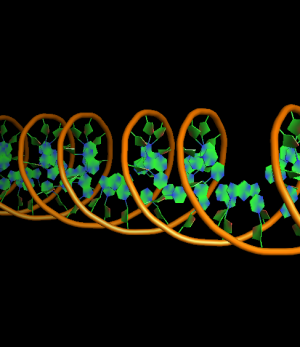
set cartoon_ring_mode, 2 # filled rings extending to inside edge of bonds
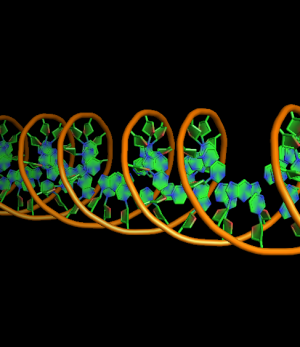
set cartoon_ring_mode, 3 # filled rings with bonds as thicker lines
cartoon ladder modes
set cartoon_ladder_mode, 0 # no ladder
set cartoon_ladder_mode, 1 # with ladder, as stick (if ring mode 0) or link to ring (if rings)
note that the visibility of the ladder sticks depends on ring mode >0, ring finder >0, nucleic acid mode = 0
all with: cartoon_ring_mode,3 cartoon_nucleic_acid_mode,0 cartoon_ring_finder,1
cartoon nucleic acid modes
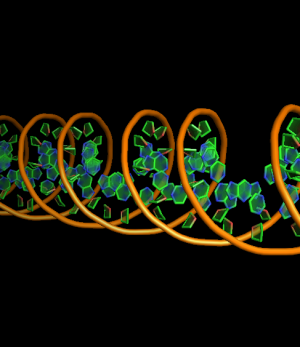
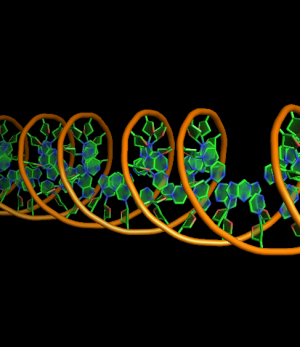
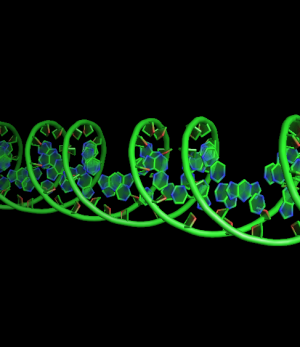
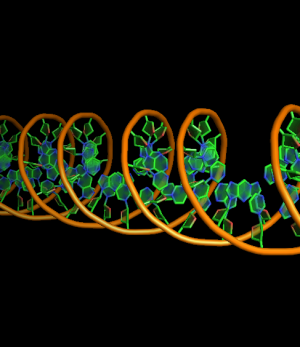
set cartoon_nucleic_acid_mode, 0 # backbone follow phosphates (red)
set cartoon_nucleic_acid_mode, 1 # backbone follows C4' of ribose (green)
all with: cartoon_ladder_mode,0 cartoon_ring_mode,3 cartoon_ring_finder,1
cartoon ring finder
set cartoon_ring_finder, 0 # no ribose, base (or ladder)
set cartoon_ring_finder, 1 # ribose and base ring
set cartoon_ring_finder, 2 # base ring only
all with: cartoon_ladder_mode,1 cartoon_ring_mode,3 cartoon_nucleic_acid_mode,0