ColorByRMSD
Jump to navigation
Jump to search
The printable version is no longer supported and may have rendering errors. Please update your browser bookmarks and please use the default browser print function instead.
| Type | Python Module |
|---|---|
| Download | colorbyrmsd.py |
| Author(s) | Shivender Shandilya, Jason Vertrees, Thomas Holder |
| License | BSD-2-Clause |
| This code has been put under version control in the project Pymol-script-repo | |
Introduction
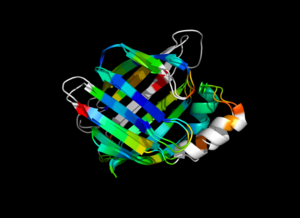
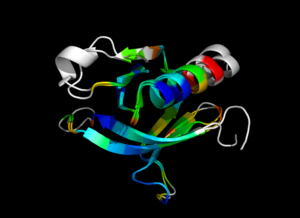
This script allows you to color two structures by Root Mean Square Deviation (RMSD). The distances between aligned C-alpha atom pairs are stored as B-factors of these residues, which are colored by a color spectrum, with blue specifying the minimum pairwise RMSD and red indicating the maximum. Unaligned residues are colored gray.
Usage
colorbyrmsd mobile, target [, doAlign [, doPretty [, guide [, method ]]]]
Arguments
- mobile = string: atom selection for mobile atoms
- target = string: atom selection for target atoms
- doAlign = 0 or 1: Superpose selections before calculating distances {default: 1}
- doPretty = 0 or 1: Show nice representation and colors {default: 1}
- guide = 0 or 1: Only use C-alpha atoms {default: 1}
- method = align or super: Method to match atoms {default: super}
Examples
# example #1
colorbyrmsd 1cbs, 1hmt, doAlign=1, doPretty=1
# example #2
colorbyrmsd 1eaz, 1fao, doAlign=1, doPretty=1