Difference between revisions of "Ccp4 contact"
(added a new script to parse and select CONTACT output) |
m (Authors corrected.) |
||
| (7 intermediate revisions by 2 users not shown) | |||
| Line 1: | Line 1: | ||
| − | [[ | + | {{Infobox script-repo |
| + | |type = script | ||
| + | |filename = ccp4_contact.py | ||
| + | |author = [[User:Dalyte|Gerhard Reitmayr and Dalia Daujotyte]] | ||
| + | |license = GPL | ||
| + | }} | ||
== Overview == | == Overview == | ||
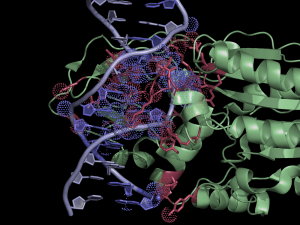
| + | [[File:HhaExample.png|thumb|300px|right|Interface residues (at cutoff <4A) in the 2c7r.pdb were found using NCONT, but similar results can be obtained using this script and CONTACT output. Usage of ccp4_contact script in PyMOL allows easy selection of residues and atoms listed in the output file. Interacting protein and DNA residues are colored in red and slate, respectively. Atoms in contact are shown in dots.]] | ||
The script selects residues and atoms from the list of the contacts found by CONTACT from CCP4 Program Suite (CONTACT analyses contacts between subsets of atoms in a PDB file). | The script selects residues and atoms from the list of the contacts found by CONTACT from CCP4 Program Suite (CONTACT analyses contacts between subsets of atoms in a PDB file). | ||
| − | First, we run CONTACT on our pdb file to find interface residues. Then by using the | + | First, we run CONTACT on our pdb file to find interface residues. Then by using the ccp4_contact script in PyMOL we separately select residues and atoms listed in the output file. This generates two selections (atoms and residues) for each interacting chain, allowing quick manipulation of (sometimes) extensive lists in CONTACT log file. |
== Usage == | == Usage == | ||
| − | + | ccp4_contact( contactsfile, selName1 = "source", selName2 = "target" ) | |
| − | == | + | == Example 1 == |
| − | + | First use CONTACT to find interface residues/atoms in the pdb file. Once you have the log file proceed to PyMOL. | |
| − | + | Make sure you import the ccp4_contact script first. | |
| − | First use CONTACT to find interface residues/atoms in the pdb file. Once you have | ||
| − | Make sure you | ||
fetch 2c7r | fetch 2c7r | ||
| − | + | ccp4_contact 2c7r.contact, selName1=prot, selName2=dna | |
[[File:HhaI20example.png|thumb|300px|right|Quick and easy selection of interacting residues and atoms listed in the CONTACT log file. Protein and DNA residues are colored in red and slate, respectively. Atoms in contact are shown in dots.]] | [[File:HhaI20example.png|thumb|300px|right|Quick and easy selection of interacting residues and atoms listed in the CONTACT log file. Protein and DNA residues are colored in red and slate, respectively. Atoms in contact are shown in dots.]] | ||
| − | + | {{Template:PymolScriptRepoDownload|examples/ccp4_contact_1.pml}} | |
| − | + | <include src="https://raw.github.com/Pymol-Scripts/Pymol-script-repo/master/examples/ccp4_contact_1.pml" highlight="python" /> | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | == Getting a CONTACT file == | |
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | + | === Install CCP4 - for Linux === | |
| − | < | + | Goto: http://www.ccp4.ac.uk/download.php <br> |
| + | Click: automated Downloads Pages <br> | ||
| + | Select: Linux, generic linux (x86) <br> | ||
| + | Select: Customized installation <br> | ||
| + | Select: Only CCP4 Program Suite, Executables -> Continue <br> | ||
| + | No additional packages -> Continue <br> | ||
| + | Download <br> | ||
| − | == | + | Extract for example to: '''/home/YOU/Software/CCP'''4 <br> |
| + | Then run: <br> | ||
| + | <syntaxhighlight lang="bash"> | ||
| + | $ /home/YOU/Software/CCP4/install.sh | ||
| + | </syntaxhighlight> | ||
| + | write yes, read agreement, push y to agree license <br> | ||
| + | For sourcing scripts, say yes. <br> | ||
| + | See the changes to your environmental virables: <br> | ||
| + | <syntaxhighlight lang="bash"> | ||
| + | $ less ~/.bashrc | ||
| + | </syntaxhighlight> | ||
| − | + | === Use of CONTACT - for Linux === | |
| + | See here for the CONTACT program and options: http://www.ccp4.ac.uk/html/contact.html <br> | ||
| + | Locate the pdb, and now run in terminal: <br> | ||
| + | <syntaxhighlight lang="bash"> | ||
| + | $ contact XYZIN 2c7r.pdb >> 2c7r.contact << eof (#press enter) | ||
| + | > MODE ISUB (#press enter) | ||
| + | > ATYPE NON-CARBON (#press enter) | ||
| + | > eof (#press enter, and now the program runs, and shell saves to 2c7r.contact) | ||
| + | </syntaxhighlight> | ||
| − | [[Category:Script_Library]] [[Category:ThirdParty Scripts]] [[Category:Structural Biology Scripts]] | + | [[Category:Script_Library]] |
| + | [[Category:ThirdParty Scripts]] | ||
| + | [[Category:Structural Biology Scripts]] | ||
| + | [[Category:Pymol-script-repo]] | ||
Latest revision as of 18:15, 13 February 2012
| Type | Python Script |
|---|---|
| Download | ccp4_contact.py |
| Author(s) | Gerhard Reitmayr and Dalia Daujotyte |
| License | GPL |
| This code has been put under version control in the project Pymol-script-repo | |
Overview
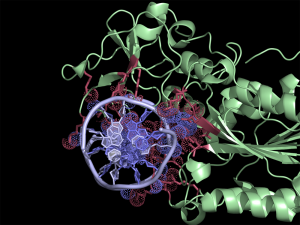
The script selects residues and atoms from the list of the contacts found by CONTACT from CCP4 Program Suite (CONTACT analyses contacts between subsets of atoms in a PDB file). First, we run CONTACT on our pdb file to find interface residues. Then by using the ccp4_contact script in PyMOL we separately select residues and atoms listed in the output file. This generates two selections (atoms and residues) for each interacting chain, allowing quick manipulation of (sometimes) extensive lists in CONTACT log file.
Usage
ccp4_contact( contactsfile, selName1 = "source", selName2 = "target" )
Example 1
First use CONTACT to find interface residues/atoms in the pdb file. Once you have the log file proceed to PyMOL. Make sure you import the ccp4_contact script first.
fetch 2c7r ccp4_contact 2c7r.contact, selName1=prot, selName2=dna
| Download: examples/ccp4_contact_1.pml | |
| This code has been put under version control in the project Pymol-script-repo | |
reinitialize
import ccp4_contact
fetch 2c7r, async=0
remove solvent
show_as cartoon, 2c7r
python
if 'PYMOL_GIT_MOD' in os.environ:
example_dir = os.path.join(os.path.split(os.environ['PYMOL_GIT_MOD'])[0],"files_for_examples")
contactfile = os.path.join(example_dir,"2c7r.contact")
else:
contactfile = "2c7r.contact"
python end
select ligands, organic
select prot, chain A
select ssDNAa, chain C
select ssDNAb, chain D
select dsDNA, chain C+D
ccp4_contact.ccp4_contact(contactfile, selName1="prot", selName2="dsDNA")
# See here to represent nuc acids
#http://www.pymolwiki.org/index.php/Examples_of_nucleic_acid_cartoons
set cartoon_ring_mode, 3
set cartoon_ring_finder, 1
color slate, dsDNA and elem C
show sticks, prot_res and prot
color raspberry, prot_res and prot
show dots, prot_atom and prot
orient prot_res
ray
Getting a CONTACT file
Install CCP4 - for Linux
Goto: http://www.ccp4.ac.uk/download.php
Click: automated Downloads Pages
Select: Linux, generic linux (x86)
Select: Customized installation
Select: Only CCP4 Program Suite, Executables -> Continue
No additional packages -> Continue
Download
Extract for example to: /home/YOU/Software/CCP4
Then run:
$ /home/YOU/Software/CCP4/install.sh
write yes, read agreement, push y to agree license
For sourcing scripts, say yes.
See the changes to your environmental virables:
$ less ~/.bashrc
Use of CONTACT - for Linux
See here for the CONTACT program and options: http://www.ccp4.ac.uk/html/contact.html
Locate the pdb, and now run in terminal:
$ contact XYZIN 2c7r.pdb >> 2c7r.contact << eof (#press enter)
> MODE ISUB (#press enter)
> ATYPE NON-CARBON (#press enter)
> eof (#press enter, and now the program runs, and shell saves to 2c7r.contact)