Annotation wizard
Jump to navigation
Jump to search
The printable version is no longer supported and may have rendering errors. Please update your browser bookmarks and please use the default browser print function instead.
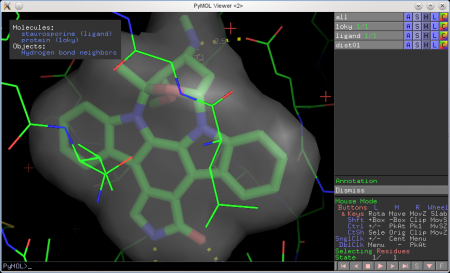
Here's an example of using the annotation wizard (see top-left corner):
import pymol
# fetch a protein and setup the view
cmd.fetch("1oky", async=0)
cmd.extract("ligand", "resn STU")
cmd.show_as("sticks", "resn STU")
cmd.show("surface", "ligand")
cmd.flag("ignore", "not rep surface")
cmd.set("surface_color", "grey")
cmd.set("transparency", 0.3)
cmd.distance("(ligand)", "(poly)", quiet=1, mode=2, label=1)
# turn on the annotation wizard + prompt
cmd.wizard("annotation")
cmd.set("wizard_prompt_mode", 1)
pymol.session.annotation = {}
state_dict = {1: ['\\999Molecules:',' \\459staurosporine (ligand)',' \\459protein (1oky)','\\999Objects:',' \\459Hydrogen bond neighbors',]}
pymol.session.annotation["1oky"] = state_dict
The following source shows you how to use the annotation wizard for multiple objects.
import pymol
# fetch a protein and setup the view
cmd.fetch("1oky", async=0)
cmd.extract("ligand", "resn STU")
cmd.show_as("sticks", "resn STU")
cmd.show("surface", "ligand")
cmd.flag("ignore", "not rep surface")
cmd.set("surface_color", "grey")
cmd.set("transparency", 0.3)
cmd.distance("(ligand)", "(poly)", quiet=1, mode=2, label=1)
# turn on the annotation wizard + prompt
cmd.wizard("annotation")
cmd.set("wizard_prompt_mode", 1)
pymol.session.annotation = {}
# using the annotation wizard for multiple objects:
prot_dict = { 1: ['\\999Protein:',' \\459protein (1oky)',]}
lig_dict = { 1: ['\\999Ligand:',' \\459staurosporine (ligand)',]}
dist_dict = { 1: ['\\999Objects: ',' \\459Hydrogen bond neighbors',]}
pymol.session.annotation["1oky"] = prot_dict
pymol.session.annotation["ligand"] = lig_dict
pymol.session.annotation["dist01"] = dist_dict