Difference between revisions of "Main Page"
(new script: uniprot_features) |
|||
| (3 intermediate revisions by 3 users not shown) | |||
| Line 26: | Line 26: | ||
|- | |- | ||
! New Script | ! New Script | ||
| − | | [[dehydron]] A plugin to calculate dehydrons and display them onto the protein structure. | + | | [[uniprot_features]] makes named selections for sequence annotations from uniprot |
| − | A dehydron is a main chain hydrogen bond incompletely shielded from water attack. | + | |- |
| + | ! New Plugin | ||
| + | | [[Gyration_tensor]] Calculates chain-wise gyration tensor of a protein. | ||
| + | |- | ||
| + | ! New Script | ||
| + | | [[set_phipsi]] can set phi/psi angles for all residues in a selection | ||
| + | |- | ||
| + | ! New Script | ||
| + | | [[dehydron]] A plugin to calculate dehydrons and display them onto the protein structure. A dehydron is a main chain hydrogen bond incompletely shielded from water attack. | ||
|- | |- | ||
! New Script | ! New Script | ||
| − | | [[pymol2glmol]] is script to export a scene in pymol to a webpage for GLmol. GLmol is a molecular viewer for Web browsers written in WebGL/Javascript. | + | | [[pymol2glmol]] is script to export a scene in pymol to a webpage for GLmol. GLmol is a molecular viewer for Web browsers written in WebGL/Javascript. Like web Jmol, but MUCH faster. Try it out! |
| − | Like web Jmol, but MUCH faster. Try it out! | ||
|- | |- | ||
! New Script | ! New Script | ||
Revision as of 06:35, 13 February 2012
| The community-run support site for the PyMOL molecular viewer. |
| New accounts: email jason (dot) vertrees (@) gmail dot com |
| Tutorials | Table of Contents | Commands |
| Script Library | Plugins | FAQ |
| Gallery | Covers | PyMOL Cheat Sheet (PDF) | GoogleSearch |
|
|
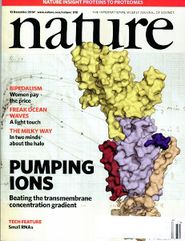 A Random PyMOL-generated Cover. See Covers.
|