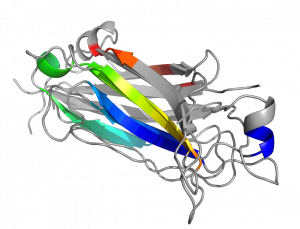
This script will align and color the paired secondary structures of the two proteins a similar rainbow color.
<source lang="python">
from pymol import cmd, util
def highlight_aligned_ss(obj1,obj2,transform=1,quiet=1):
"""
DESCRIPTION
Aligns two structures and colors their matching
secondary structure elements with a matching
rainbow colorscheme.
USAGE
highlight_aligned_ss obj1, obj2
If transform=0 then the proteins are not
moved after alignment.
EXAMPLES
highlight_aligned_ss 1cll, 1ggz
highlight_aligned_ss 1rlw, 1byn and state 1
SEE ALSO
align
JV 3-2-11
"""
if not cmd.count_atoms(obj1):
print "Error: Object 1 needs at least a few atoms to align."
return None
if not cmd.count_atoms(obj2):
print "Error: Object 2 needs at least a few atoms to align."
return None
# align them
uAln = cmd.get_unused_name("aln")
cmd.align(obj1,obj2,object=uAln,transform=int(transform))
..→